2026-06-02 23:59:59
SecurityMetrics has won an award for their tool, Shopping Cart Monitor (SCM) which helps SMBs strengthen their cybersecurity posture and defend against e-commerce threats.
2026-06-02 23:54:51
to be deleted
2026-06-02 22:40:31
PC Workman is a Windows system-monitoring and optimization tool that uses a local AI assistant, hck_GPT, to explain system behavior in plain language rather than simply displaying hardware statistics. By building a personalized baseline for each machine, the software can identify unusual activity, answer performance-related questions, and recommend optimizations based on actual usage patterns. The project currently has a Proof of Usefulness score of 58 and has demonstrated strong build-in-public engagement, with user feedback directly shaping new features
2026-06-02 22:15:55
Enterprise legal teams are signing AI agreements they don't fully understand. Enterprise engineering teams are building on top of those agreements without reading them. The result is a compliance gap that won't surface until it's too late.
I've reviewed a lot of enterprise vendor agreements in my consulting work. SaaS contracts, cloud infrastructure MSAs, data processing addendums. The language is usually dense, the protections usually narrower than the sales pitch, and the gaps usually invisible until an audit or an incident forces everyone to look.
The AI agreements I've been reviewing over the past eighteen months are in a different category entirely. Not because the lawyers are less skilled — they're not — but because the underlying technology is complex enough that the legal language routinely fails to capture what's actually happening at the infrastructure level.
The specific clause I keep seeing misunderstood: "We do not train our models on your data."
This clause is real. It's in most enterprise AI agreements. It's also much narrower than almost every enterprise buyer assumes it to be.
Let me break down exactly what it covers, what it doesn't, and what the actual risk surface looks like for companies relying on it as a primary data protection mechanism.
When an AI provider writes "we do not train on customer data," they are making a specific, bounded commitment: the text you send through their API will not be used to update the weights of their foundation models.
That's it. That's the commitment.
It does not mean:
I'm not describing theoretical risks. These are documented behaviors in standard enterprise AI agreements, if you read the full data processing addendum rather than the marketing summary.
Layer 1: Inference Logging
Most enterprise AI providers log API requests for abuse detection, rate limiting, and service reliability monitoring. The retention period varies — it's typically documented in the DPA, often 30 days, sometimes longer. During that window, your compiled prompts — including the retrieved proprietary context from your RAG pipeline — exist on the provider's infrastructure.
"Zero training" doesn't touch this. These are operational logs, not training data.
Layer 2: Prompt Caching
Several major providers have introduced prompt caching as a latency optimization feature. When enabled, frequently-used prompt prefixes are stored in the provider's infrastructure to reduce repeated computation costs. For enterprise RAG pipelines where the system prompt contains proprietary context, this means your data may be cached on external infrastructure for the duration of the cache TTL.
Read your provider's documentation on whether prompt caching is opt-in or opt-out for enterprise tiers. The answer will vary, and the default may not be what you assumed.
Layer 3: Subprocessor Infrastructure
Your enterprise agreement is with the AI provider. But that provider's inference infrastructure runs on hyperscaler cloud services — AWS, GCP, Azure — under the provider's cloud agreements, not yours. Your data processing addendum with the AI provider may have strong protections. The subprocessor chain beneath it is governed by agreements you've never seen.
This matters particularly for GDPR compliance, where Article 28 requires documented subprocessor chains with equivalent protections. "Our cloud provider also has a strong DPA" is not the same as having reviewed it.
Layer 4: Jurisdictional Exposure
If the AI provider is a US-based company, your data — regardless of where their servers are physically located — is potentially subject to US legal process under the Stored Communications Act and related statutes. If your enterprise handles data subject to GDPR, you've now got a potential conflict between your data residency obligations and the jurisdictional reach of your AI vendor's legal exposure.
This isn't hypothetical. It's the same issue that forced the EU's invalidation of Privacy Shield in 2020 and continues to create compliance headaches for multinational enterprises.
Let me be specific about three frameworks I see most frequently in enterprise AI deployments.
GDPR (General Data Protection Regulation)
GDPR doesn't prohibit sending personal data to third-party processors. It requires that you have a lawful basis for the transfer, a data processing agreement with the processor, documented subprocessor chains, and — for transfers outside the EEA — an appropriate transfer mechanism (SCCs, adequacy decision, etc.).
A "zero training" clause is not a transfer mechanism. It's a use restriction. These are different things. If your enterprise processes EU personal data through an external AI API, you need the full legal infrastructure, not just a favorable marketing clause.
SOC 2 Type II
SOC 2 audits your internal controls. It doesn't audit your vendors. Having an AI vendor with their own SOC 2 report is good, but it doesn't substitute for your own access controls, data classification, and vendor management processes. In the post-incident reviews I've participated in, "the vendor has SOC 2" is consistently one of the weaker defenses in an audit finding.
HIPAA
If you're in healthcare and you're sending any PHI-adjacent data through an external AI API — even indirectly through a RAG pipeline that indexes patient records — you need a signed BAA with the provider, and the BAA needs to be specific about the AI use case. Generic cloud infrastructure BAAs don't cover LLM inference use cases. This gap has already produced compliance findings at several healthcare organizations I'm aware of.
When I review AI vendor agreements with enterprise clients, I'm looking for answers to these specific questions. Most enterprise buyers have never asked them.
1. What is the full data retention schedule across all pipeline layers? Not just "we don't train." What is retained, where, for how long, and under what deletion policy? Get the answer in the DPA, not the sales deck.
2. What is the complete subprocessor list, and are their DPAs equivalent? Request the current subprocessor list. It should be in the agreement or available on demand. Verify that subprocessors have equivalent data protection commitments.
3. What is the default state of prompt caching, logging, and data residency for your tier? "Enterprise" tiers often have different defaults than standard tiers. Confirm the specific configuration that applies to your agreement, in writing.
4. What is the provider's legal response protocol for government data requests? How does the provider handle subpoenas, national security letters, and foreign government requests? Do they commit to notifying customers before complying with legal process, to the extent legally permitted? What jurisdiction governs?
5. What is the incident response SLA, and what data does it cover? If the provider has a security incident that exposes your prompts from the inference logging layer, what is their notification obligation and timeline? "Zero training data" is irrelevant if the incident involves inference logs.
I want to be precise about something: the legal issues I've described above are a symptom of an architectural decision, not a standalone problem.
When your AI pipeline sends proprietary data to an external inference endpoint, you've created a legal exposure because you've created an architectural exposure. The two are inseparable. Stronger contract language reduces the legal risk at the margins. It doesn't change the underlying data flow.
The enterprises I've seen handle this correctly have approached it as an architecture problem first and a vendor management problem second. The design goal is to keep the data and the inference engine in the same security and legal perimeter, so the vendor agreement question becomes much simpler: what does the vendor have access to in the first place?
Self-hosted inference — whether a custom Kubernetes deployment or a unified self-hosted platform that runs the orchestration and inference layer on your own infrastructure — doesn't eliminate vendor relationships, but it fundamentally changes what those relationships cover. Your vendor agreement governs software licensing and support. Your data never leaves your environment in the first place, so the DPA questions about inference logging, prompt caching, and subprocessor chains become irrelevant.
That's a much cleaner compliance posture to maintain and audit.
If you're an enterprise that has deployed or is evaluating external AI API integrations, here's where to start:
Pull the current data processing addendum for every AI vendor in your stack. Not the master service agreement — the DPA. Read the retention schedules, the subprocessor list, and the security incident notification clauses specifically.
Then map that against the actual data flowing through your AI pipeline. What data is being retrieved and compiled into prompts? How is it classified? Does your current DPA coverage match the sensitivity of that data?
The gap between those two answers is your current compliance exposure. Most enterprises I've worked with have a larger gap than they realize, because the "zero training" clause felt sufficient and nobody looked further.
It isn't sufficient. Look further.
Tags: #enterprise-ai #gdpr #compliance #data-privacy #ai-security #llm #legal-tech #rag #vendor-management #global-ip
\
2026-06-02 22:01:16
How are you, hacker?
🪐Want to know what's trending right now?:
The Techbeat by HackerNoon has got you covered with fresh content from our trending stories of the day! Set email preference here.
## Why macOS Is Underrepresented in Public AI Research Datasets  By @macpaw [ 5 Min read ]
MacPaw Research explains why macOS is severely underrepresented in public AI datasets and introduces GUIrilla, a framework for scalable Mac UI exploration. Read More.
By @macpaw [ 5 Min read ]
MacPaw Research explains why macOS is severely underrepresented in public AI datasets and introduces GUIrilla, a framework for scalable Mac UI exploration. Read More.

By @playerzero [ 8 Min read ] PlayerZero explains how predictive software quality helps enterprises prevent defects, reduce firefighting, and scale reliable software development with AI. Read More.

By @scylladb [ 4 Min read ] New research reveals why companies ignore database risks until costs, latency, or outages force migrations—and what warning signs to watch. Read More.

By @apilayer [ 4 Min read ] Discover 7 of the best SERP APIs for SEO and market research in 2026. Read More.

By @qatech [ 7 Min read ] Learn how AI-generated code is shifting software QA toward continuous verification, agentic testing, and context-aware product quality systems. Read More.

By @kumar96 [ 7 Min read ] Creating third-party integration is not a problem in OutSystems. It talks about web services, OAuth, and other security in APIs Read More.

By @webintelligencehub [ 7 Min read ] IP pool size is a marketing number. Learn which metrics matter when evaluating proxy providers, and what to look for on their websites before you spend a dollar Read More.

By @akashi-ghost [ 11 Min read ] AI is not alien or fake intelligence. It is accumulated human thought, culture, code, bias, and memory reflected back through machines. Read More.

By @bzimbelman [ 11 Min read ] AI coding agents need more than local shells. Shared pools, reservations, and project definitions make enterprise-scale orchestration possible. Read More.

By @oleksiijko [ 14 Min read ] Open-source local-first memory for Claude Code, Cursor, Codex via MCP. 94.5% LoCoMo recall, 70ms p50, no API keys. Five techniques explained with code. Read More.

By @pauliusuza [ 5 Min read ] Most AI products lose users after day two. Here are the four failure modes that decide whether they come back — and the design patterns that fix each one. Read More.

By @nazar-kozak [ 10 Min read ] This article explores how Claude Code memory layers, task-scoped sessions, and context management can improve AI-assisted software development workflows. Read More.

By @zbruceli [ 21 Min read ] Inside the biological tricks that let three-pound brains outlearn trillion-parameter machines — and the radical new architectures trying to close the gap Read More.

By @gtechguide123 [ 9 Min read ] A comprehensive guide to AI tech stacks, covering data infrastructure, machine learning frameworks, MLOps, and Generative AI development. Read More.

By @speechmatics [ 8 Min read ] A comparison of seven voice agent testing platforms, covering audio quality, simulation, observability, compliance, and regression testing. Read More.

By @kilocode [ 6 Min read ] This article explores how AI developer tools are moving beyond the IDE to reduce context switching across Slack, cloud agents, and coding workflows. Read More.

By @efimovov_5guqm5 [ 7 Min read ] Skills extend Claude Code with reusable slash commands — but auto-invocation depends on description quality, and silent failures are common. Read More.

By @huckler [ 7 Min read ] My system monitor with its own 9-layer AI between €3.8/h retail shifts. 55 GitHub stars, 260 downloads, 10 months solo. Here's everything it does. PC Workman. Read More.

By @vladlensk1y [ 4 Min read ] How I use AGENTS.md and a GitHub Action to catch low-effort AI-generated PRs before wasting maintainer review time. Read More.

By @sodax [ 4 Min read ]
Wrapped Bitcoin was a workaround. The technology’s ready for native BTC in DeFi money markets. The SODAX SDK is the operational version. Read More.
🧑💻 What happened in your world this week? It's been said that writing can help consolidate technical knowledge, establish credibility, and contribute to emerging community standards. Feeling stuck? We got you covered ⬇️⬇️⬇️
ANSWER THESE GREATEST INTERVIEW QUESTIONS OF ALL TIME
We hope you enjoy this worth of free reading material. Feel free to forward this email to a nerdy friend who'll love you for it.
See you on Planet Internet! With love,
The HackerNoon Team ✌️
.gif)
2026-06-02 22:00:49
“If you always do what you always did, you will always get what you always got.” ― Albert Einstein
 I had a lot of questions on my quest to understand how Blockchain works. The important one was “How do I build applications on it?”. It took a few weeks of digging up, reading and experimenting to finally get it. I couldn’t find a short but comprehensive guide. Now, that I have some decent understanding, I thought of writing one that could help others. This is a light speed guide, I have kept only the important parts in order to reduce the learning curve.
I had a lot of questions on my quest to understand how Blockchain works. The important one was “How do I build applications on it?”. It took a few weeks of digging up, reading and experimenting to finally get it. I couldn’t find a short but comprehensive guide. Now, that I have some decent understanding, I thought of writing one that could help others. This is a light speed guide, I have kept only the important parts in order to reduce the learning curve.
 A reflection on why true automation starts with human thinking, not technology. Systems only work as clearly as the minds that design them.
A reflection on why true automation starts with human thinking, not technology. Systems only work as clearly as the minds that design them.

 In this article, we will learn and break down what AI agents are, how they think, and how they are shaping the tech world and becoming part of our daily lives.
In this article, we will learn and break down what AI agents are, how they think, and how they are shaping the tech world and becoming part of our daily lives.
 What do all recent super powerful image models like DALLE, Imagen, or Midjourney have in common? Other than their high computing costs, huge training time, and shared hype, they are all based on the same mechanism: diffusion.
What do all recent super powerful image models like DALLE, Imagen, or Midjourney have in common? Other than their high computing costs, huge training time, and shared hype, they are all based on the same mechanism: diffusion.
 In the present dynamic and competitive business environment, product managers play a crucial role in pushing innovation…
In the present dynamic and competitive business environment, product managers play a crucial role in pushing innovation…
 We skipped last year, but this year SparkLabs Group is back with our fourth
rankings and report of the Top Ten Startup Ecosystems in the World.
We skipped last year, but this year SparkLabs Group is back with our fourth
rankings and report of the Top Ten Startup Ecosystems in the World.
 Two strategies to integrate UX research in banking and retail, overcoming challenges to enhance product discovery and drive better results
Two strategies to integrate UX research in banking and retail, overcoming challenges to enhance product discovery and drive better results
 How we transformed innovation culture by balancing creativity with structure, overcoming bureaucracy, and fostering growth—insights from hands-on experience.
How we transformed innovation culture by balancing creativity with structure, overcoming bureaucracy, and fostering growth—insights from hands-on experience.
 This Ethereum DEX Will Cover Gas for Your Limit Orders
This Ethereum DEX Will Cover Gas for Your Limit Orders
 I love Apple. Really. Seriously. Love, Apple products. I currently use a full Apple line up; MacBook Air 11' and iPhone 6S Plus. Both products have been absolutely superb to date.
I love Apple. Really. Seriously. Love, Apple products. I currently use a full Apple line up; MacBook Air 11' and iPhone 6S Plus. Both products have been absolutely superb to date.
 Learn why the resistance we're seeing against Bitcoin today is nothing new. The same thing happened with refrigerators, airplanes, and tractors.
Learn why the resistance we're seeing against Bitcoin today is nothing new. The same thing happened with refrigerators, airplanes, and tractors.
 A tiny AI robot named Erbai staged a bizarre 'kidnapping,' convincing 12 larger robots to quit their jobs in a Shanghai showroom. A Black Mirror moment?
A tiny AI robot named Erbai staged a bizarre 'kidnapping,' convincing 12 larger robots to quit their jobs in a Shanghai showroom. A Black Mirror moment?
 Explore the limitless potential of OpenAI's GPT-4 with Kartik Khosa, as he transforms personalized study plans into diverse AI applications.
Explore the limitless potential of OpenAI's GPT-4 with Kartik Khosa, as he transforms personalized study plans into diverse AI applications.
 Upgradable NFTs will uprise as the next innovation in the non-fungible token marketplace. It will allow collectors to engage & have utility for their NFTs.
Upgradable NFTs will uprise as the next innovation in the non-fungible token marketplace. It will allow collectors to engage & have utility for their NFTs.
 Burning Man 2002. Photo: Phil Gyford
Burning Man 2002. Photo: Phil Gyford
 New research by Niv Haim et al. allows us to perform infinite video manipulations without using deep learning or datasets.
New research by Niv Haim et al. allows us to perform infinite video manipulations without using deep learning or datasets.
 After nearly 9 months of work, version 3 of “A Survey on Blockchain Interoperability: Past, Present, and Future Trends” is out. This survey depicts the state of blockchain interoperability as of March 2021, by collecting and analyzing 404 documents.
After nearly 9 months of work, version 3 of “A Survey on Blockchain Interoperability: Past, Present, and Future Trends” is out. This survey depicts the state of blockchain interoperability as of March 2021, by collecting and analyzing 404 documents.
 Innovation is a critical driver of economic growth and societal progress. But worryingly, tech innovation has been slowing for decades.
Innovation is a critical driver of economic growth and societal progress. But worryingly, tech innovation has been slowing for decades.
 TimeLens can understand the movement of the particles in-between the frames of a video to reconstruct what really happened at a speed even our eyes cannot see.
TimeLens can understand the movement of the particles in-between the frames of a video to reconstruct what really happened at a speed even our eyes cannot see.
 ChatGPT is all over the internet with people buzzing about its capabilities, but is it the future or just another gimmick? Let's find out!
ChatGPT is all over the internet with people buzzing about its capabilities, but is it the future or just another gimmick? Let's find out!
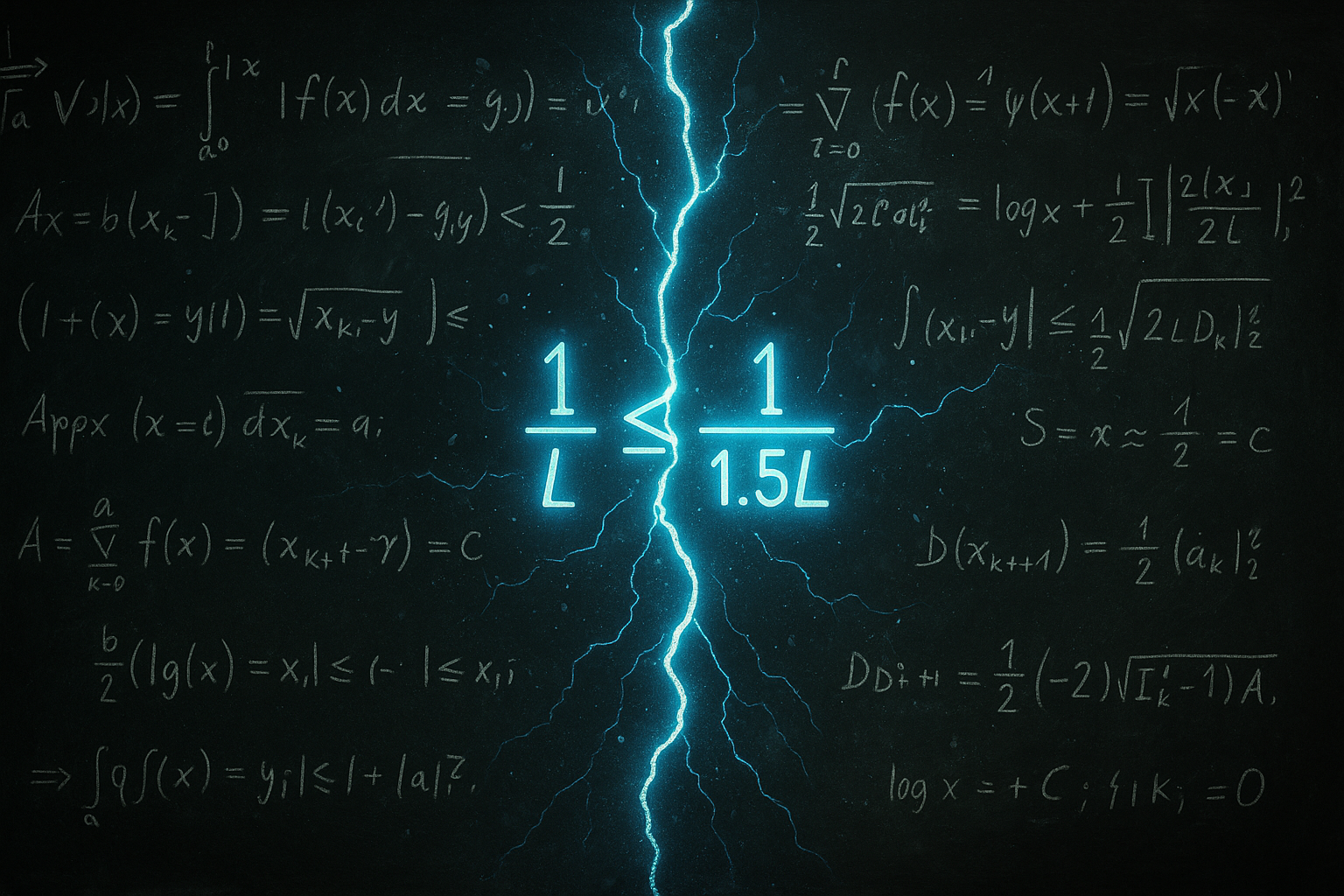 GPT-5 solved original math, ending humanity’s monopoly on discovery. Ronnie Huss breaks down what this means for AI, science, and the frontier ahead.
GPT-5 solved original math, ending humanity’s monopoly on discovery. Ronnie Huss breaks down what this means for AI, science, and the frontier ahead.
 We all from time to time come across an idea for a Startup. A company that can be a million or billion or maybe trillion-dollar worth. A company that can be next Tesla or Amazon, but sadly, 99.99999% of us don’t take the next step. That idea remains just a fragment of our memory.
We all from time to time come across an idea for a Startup. A company that can be a million or billion or maybe trillion-dollar worth. A company that can be next Tesla or Amazon, but sadly, 99.99999% of us don’t take the next step. That idea remains just a fragment of our memory.

 The current lockdown has given many people time to think about personal and professional improvements they want to pursue. We are currently watching industries being re-calibrated and some come close to the point of failure. Now is a good reminder to always keep in mind innovation and digital transformation. The risk of not doing so leaves companies and full industries at risk of massive failure.
The current lockdown has given many people time to think about personal and professional improvements they want to pursue. We are currently watching industries being re-calibrated and some come close to the point of failure. Now is a good reminder to always keep in mind innovation and digital transformation. The risk of not doing so leaves companies and full industries at risk of massive failure.
 Discovering meaningful patterns in business failures.
Discovering meaningful patterns in business failures.
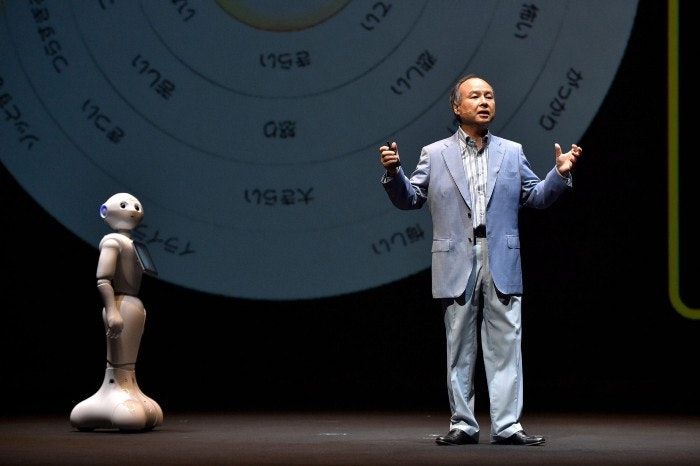
 Human civilization is fast approaching the Post Human Era or Transhuman Era where machines start to become more powerful than us. I’d say 2020 was just a taste of the overwhelming power of digital technologies such as social networks, smartphones, ubiquitous connectivity, etc. This is extraordinarily overwhelming, disastrous, scary, crazy, beautiful and awe inspiring at the same time. You feel every emotion possible to know that we are living in the future.
Human civilization is fast approaching the Post Human Era or Transhuman Era where machines start to become more powerful than us. I’d say 2020 was just a taste of the overwhelming power of digital technologies such as social networks, smartphones, ubiquitous connectivity, etc. This is extraordinarily overwhelming, disastrous, scary, crazy, beautiful and awe inspiring at the same time. You feel every emotion possible to know that we are living in the future.
 Logging into a website or service using the traditional username and password combination isn’t the best or safest way of going about it anymore.
Logging into a website or service using the traditional username and password combination isn’t the best or safest way of going about it anymore.
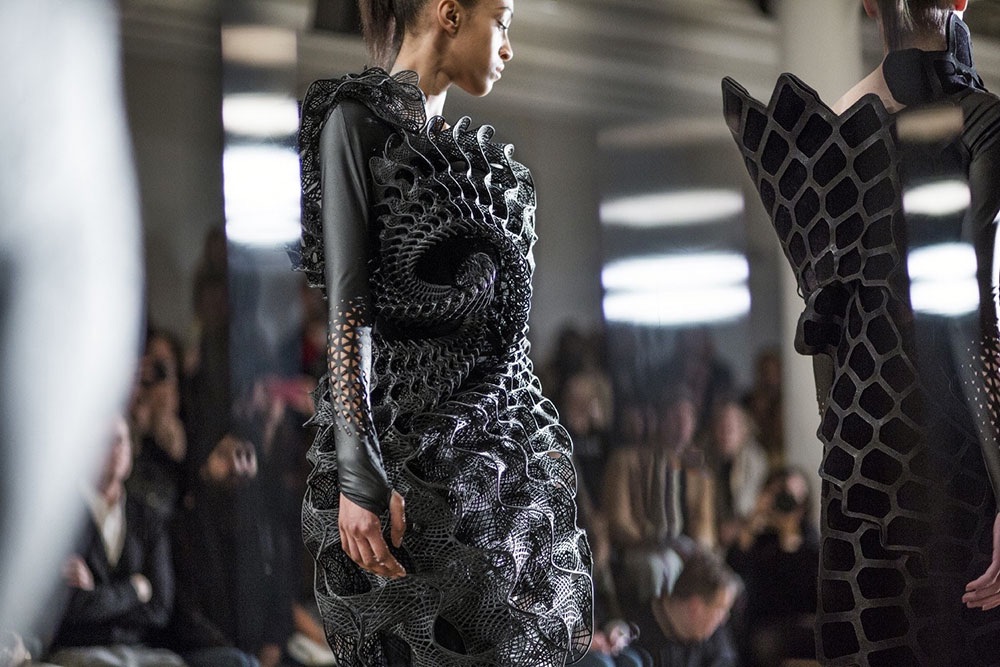 A new kind of fashion has hit the fashion industry called nanotechnology. From active membranes to heat protection, you don't want to miss out on these pieces.
A new kind of fashion has hit the fashion industry called nanotechnology. From active membranes to heat protection, you don't want to miss out on these pieces.
 Apple, today, is worth $2.74 trillion, with its relentless rally putting $ 3 trillion in view. But how did the trillion-dollar company become so successful?
Apple, today, is worth $2.74 trillion, with its relentless rally putting $ 3 trillion in view. But how did the trillion-dollar company become so successful?
 What’s the Role of Data Science in Finance?
What’s the Role of Data Science in Finance?
 A Chief Innovation Officer shares tips for growing a new breed of R&D engineering talents that are skilled in more than just conventional code using π-matrix.
A Chief Innovation Officer shares tips for growing a new breed of R&D engineering talents that are skilled in more than just conventional code using π-matrix.
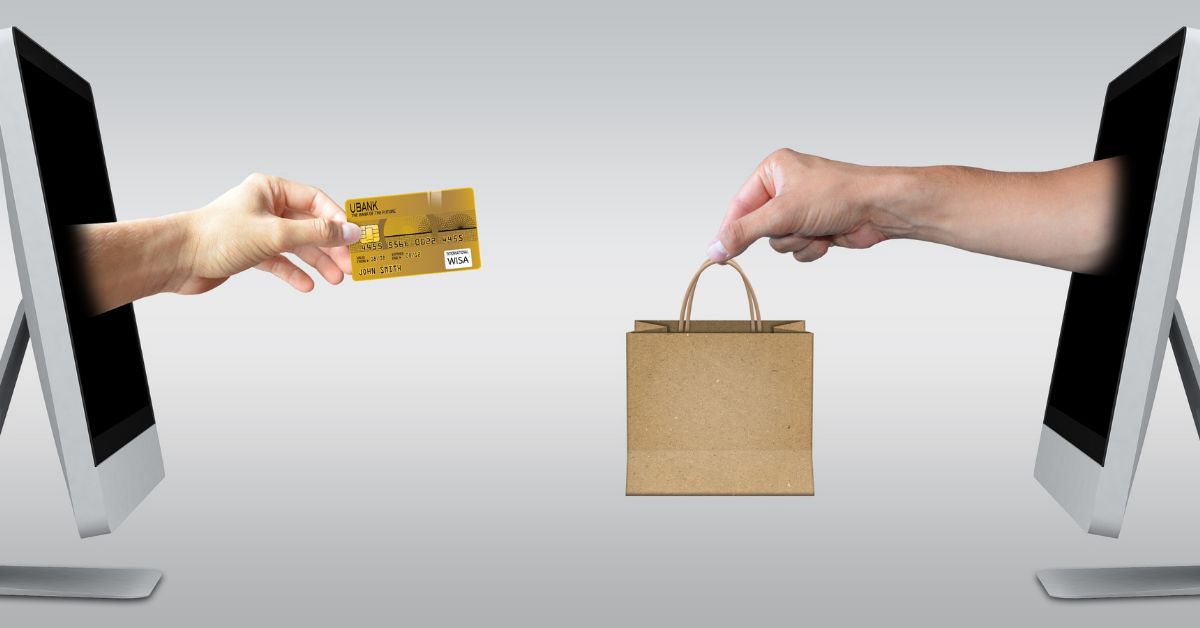 The B2C landscape is heavily affected by eCommerce and online shopping
The B2C landscape is heavily affected by eCommerce and online shopping
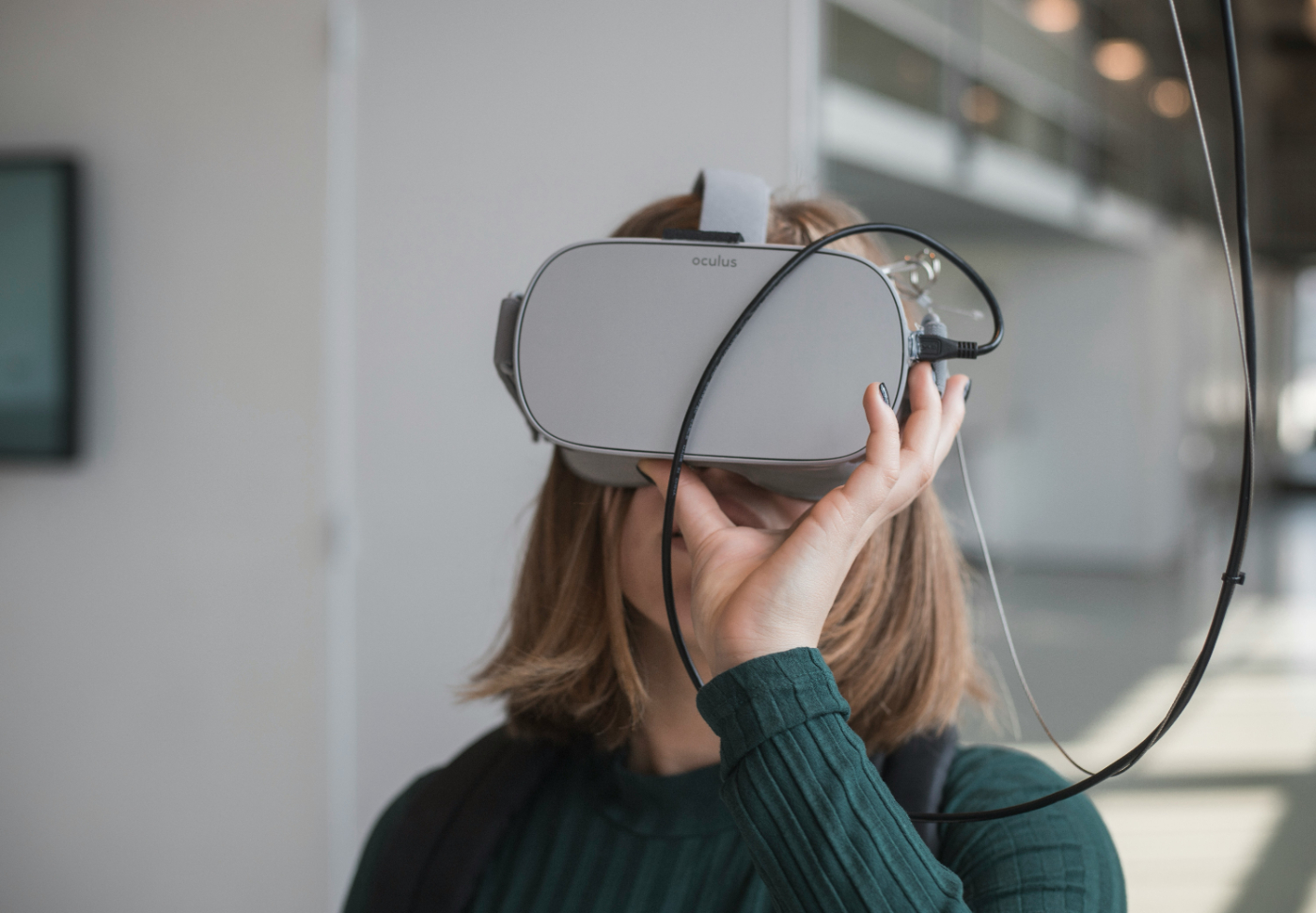 Explore the impact of low employee motivation and the role of enterprise gamification in boosting engagement, eliminating friction, and fostering creativity.
Explore the impact of low employee motivation and the role of enterprise gamification in boosting engagement, eliminating friction, and fostering creativity.

 Here is how technology is making pensions more attractive to younger generations.
Here is how technology is making pensions more attractive to younger generations.
 Jordan B. Peterson is a Canadian clinical psychologist and psychology professor at the University of Toronto who became a controversial figure in late-2016 for his critiques of political correctness. Peterson’s most recent book, 12 Rules for Life, has sold over 3 million copies worldwide. Most recently, Peterson has suffered from health issues that necessitated a year-long reprieve from the public eye.
Jordan B. Peterson is a Canadian clinical psychologist and psychology professor at the University of Toronto who became a controversial figure in late-2016 for his critiques of political correctness. Peterson’s most recent book, 12 Rules for Life, has sold over 3 million copies worldwide. Most recently, Peterson has suffered from health issues that necessitated a year-long reprieve from the public eye.
 My experience from speaking at a career event for Millennials.
My experience from speaking at a career event for Millennials.
 Amazon's Working Backwards process is well documented across the internet. But why don't more companies use this for their innovation?
Amazon's Working Backwards process is well documented across the internet. But why don't more companies use this for their innovation?
 Why change at all? It seems like a nonsensical question, doesn’t it?
Why change at all? It seems like a nonsensical question, doesn’t it?
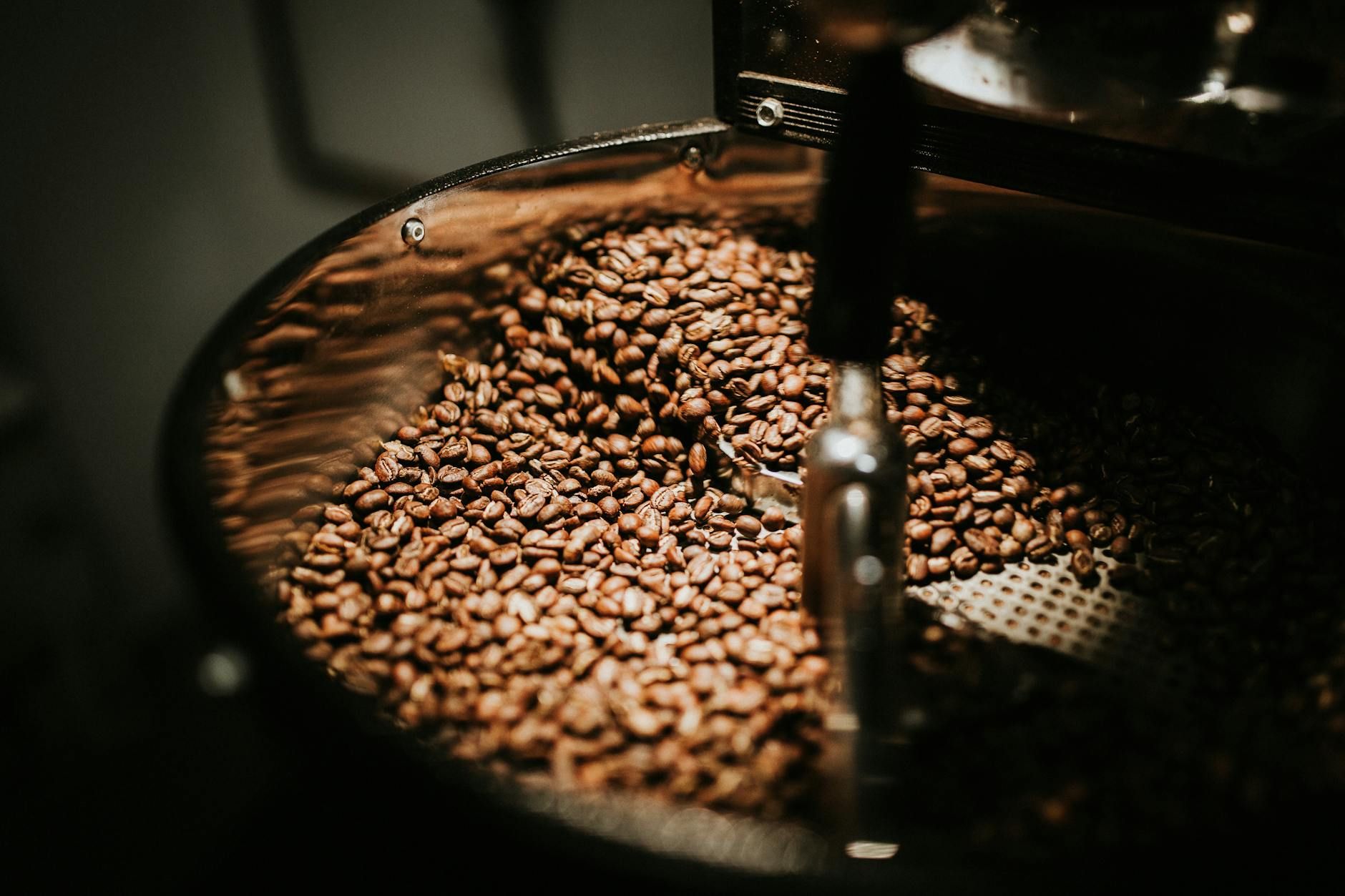 The world needs a hero to save coffee and chocolate. It’s time for Elon Musk to add BeanX to his astounding CV.
The world needs a hero to save coffee and chocolate. It’s time for Elon Musk to add BeanX to his astounding CV.
 Have you ever had an image you really liked and could only manage to find a small version of it that looked like this image above on the left? How cool would it be if you could take this image and make it twice look as good? It’s great, but what if you could make it even four or eight times more high definition? Now we’re talking, just look at that.
Have you ever had an image you really liked and could only manage to find a small version of it that looked like this image above on the left? How cool would it be if you could take this image and make it twice look as good? It’s great, but what if you could make it even four or eight times more high definition? Now we’re talking, just look at that.
 Stop being overwhelmed. Learn the neuroscience of mastery: how all genius works by compressing complexity into simple, automatic structures.
Stop being overwhelmed. Learn the neuroscience of mastery: how all genius works by compressing complexity into simple, automatic structures.
 You can read the first part of this article here. For those who for some reason don’t like to follow the links, let me remind you briefly: in the first part, we made a retrospective of fintech trends in 2020 and delved into the first 5 trends in 2021.
You can read the first part of this article here. For those who for some reason don’t like to follow the links, let me remind you briefly: in the first part, we made a retrospective of fintech trends in 2020 and delved into the first 5 trends in 2021.
 The Corporate Immune System: How To Break Through
The Corporate Immune System: How To Break Through
 Next Generation Semiconductors: Designing accelerators for advanced cryptography
Next Generation Semiconductors: Designing accelerators for advanced cryptography
 Unlock the potential of Generative AI (GenAI) with insights into its transformative impact on industries.
Unlock the potential of Generative AI (GenAI) with insights into its transformative impact on industries.
 This is not a drill. This is not the time to give up. This is not a time for excuses. This is a time for pulling out all the stops. Sounds dramatic, right?
This is not a drill. This is not the time to give up. This is not a time for excuses. This is a time for pulling out all the stops. Sounds dramatic, right?
 Drawing Bowser from Super Mario Bros in the Desmos Graphing Calculator.
Drawing Bowser from Super Mario Bros in the Desmos Graphing Calculator.
 People are always amazed when they hear that I am also a restaurant owner.
People are always amazed when they hear that I am also a restaurant owner.
 Crypto exchanges — a critical part of crypto infrastructure and much talked about in this space. Whether it’s about the huge profits being made on the big exchanges, or the hacks that happen with alarming frequency or the manner in which these exchanges are imitating, adapting and innovating upon the kinds of financial services available on Wall Street.
Crypto exchanges — a critical part of crypto infrastructure and much talked about in this space. Whether it’s about the huge profits being made on the big exchanges, or the hacks that happen with alarming frequency or the manner in which these exchanges are imitating, adapting and innovating upon the kinds of financial services available on Wall Street.
 Apple will emerge from the COVID-19 pandemic as an even more powerful and important company. Indeed, COVID-19 may prove to be Apple’s finest hour. During the first 30 days of the pandemic’s escalation in the United States, Apple stepped up in significant ways, the most notable example being the formation of a relationship with Google to contain the virus with contact-tracing technology. In addition, Apple has, among other actions:
Apple will emerge from the COVID-19 pandemic as an even more powerful and important company. Indeed, COVID-19 may prove to be Apple’s finest hour. During the first 30 days of the pandemic’s escalation in the United States, Apple stepped up in significant ways, the most notable example being the formation of a relationship with Google to contain the virus with contact-tracing technology. In addition, Apple has, among other actions:
4 Insights for Entrepreneurs](https://hackernoon.com/will-covid-19-spur-innovation-4-insights-for-entrepreneurs-us8j3yj5)
 The coronavirus pandemic has been life changing for all of us. National lockdowns, quarantines, the #stayathome campaign, remote work, interrupted travel plans, as well as struggling businesses and economies have accounted for a disruptive start to 2020. One industry that has been put to the test in the time of pandemic is healthcare.
The coronavirus pandemic has been life changing for all of us. National lockdowns, quarantines, the #stayathome campaign, remote work, interrupted travel plans, as well as struggling businesses and economies have accounted for a disruptive start to 2020. One industry that has been put to the test in the time of pandemic is healthcare.
 Interoperability is a big word with even bigger connotations for the cryptosphere. In Fact, apart from the burgeoning DeFi movement, no other industry vertical is attracting so much investment and interest.
Interoperability is a big word with even bigger connotations for the cryptosphere. In Fact, apart from the burgeoning DeFi movement, no other industry vertical is attracting so much investment and interest.
 Excellence in Software Engineering has never been a stationary destination where one can arrive sooner or later. It has always been a lifelong journey and learning process which demands consistency and commitment in order for someone to progress rapidly and to stay relevant over the next few years because of the ever-changing tech scenario. This element of uncertainty and demand for consistency has intrigued me since forever and hence compelled me to choose this a full-time career and what I’d like to do, at least for the foreseeable near future.
Excellence in Software Engineering has never been a stationary destination where one can arrive sooner or later. It has always been a lifelong journey and learning process which demands consistency and commitment in order for someone to progress rapidly and to stay relevant over the next few years because of the ever-changing tech scenario. This element of uncertainty and demand for consistency has intrigued me since forever and hence compelled me to choose this a full-time career and what I’d like to do, at least for the foreseeable near future.
 I feel disappointed; I truly tried to make it work, but the issues I found one month after buying the Samsung Galaxy Z Fold4 eventually became a dealbreaker.
I feel disappointed; I truly tried to make it work, but the issues I found one month after buying the Samsung Galaxy Z Fold4 eventually became a dealbreaker.
 In this thread, our community discusses Mojo's new product - smart contact lenses - and its potential benefits and downsides.
In this thread, our community discusses Mojo's new product - smart contact lenses - and its potential benefits and downsides.
 You need not be a natural at coming up with innovations; Use SCAMPER to generate ideas and innovate your product, service, business.
You need not be a natural at coming up with innovations; Use SCAMPER to generate ideas and innovate your product, service, business.
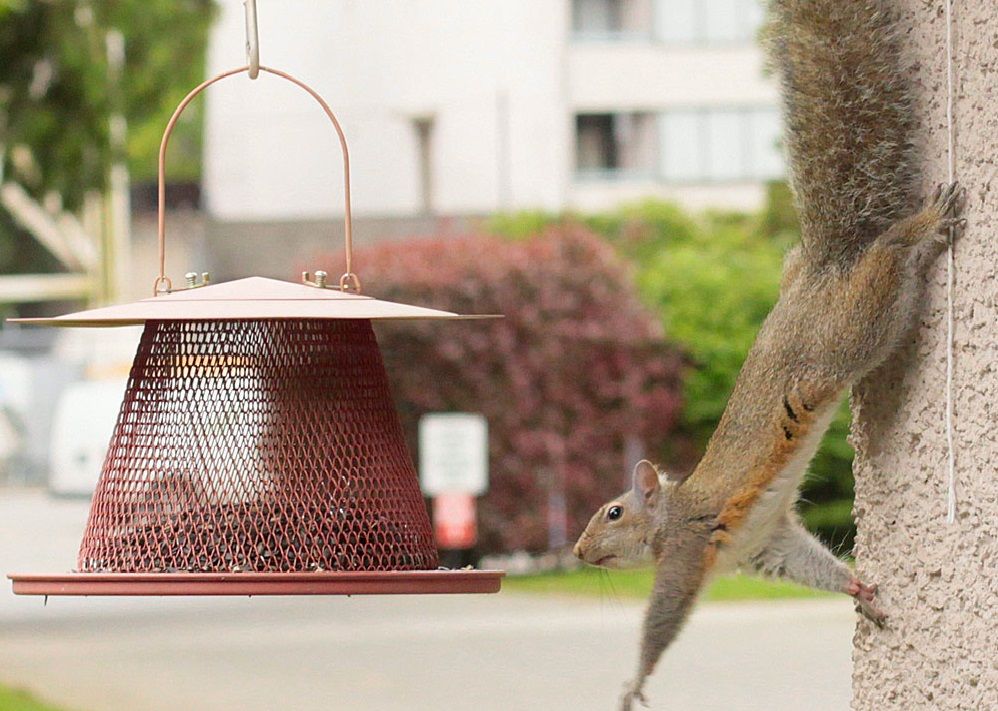 How do tech's top companies innovate at scale? It's not just Agile. According to Empowered, it's product discovery, a focus on problems, and coaching culture.
How do tech's top companies innovate at scale? It's not just Agile. According to Empowered, it's product discovery, a focus on problems, and coaching culture.
 If you think this is just like any other alarmist and opportunistic post regarding how the world is going to end with this current crisis, it is not…
If you think this is just like any other alarmist and opportunistic post regarding how the world is going to end with this current crisis, it is not…
 Explore the fascinating evolution of AI, from its humble beginnings to the cutting-edge advancements of today.
Explore the fascinating evolution of AI, from its humble beginnings to the cutting-edge advancements of today.
 Incumbent Banks and financial technology startups (Fintechs) face very different challenges as they navigate the COVID-19 crisis. Despite sharing customers and offering similar products, their business models, how they operate, their balance sheets and culture vary tremendously. Each one of these differences impacts how they will perform during and after the crisis.
Incumbent Banks and financial technology startups (Fintechs) face very different challenges as they navigate the COVID-19 crisis. Despite sharing customers and offering similar products, their business models, how they operate, their balance sheets and culture vary tremendously. Each one of these differences impacts how they will perform during and after the crisis.
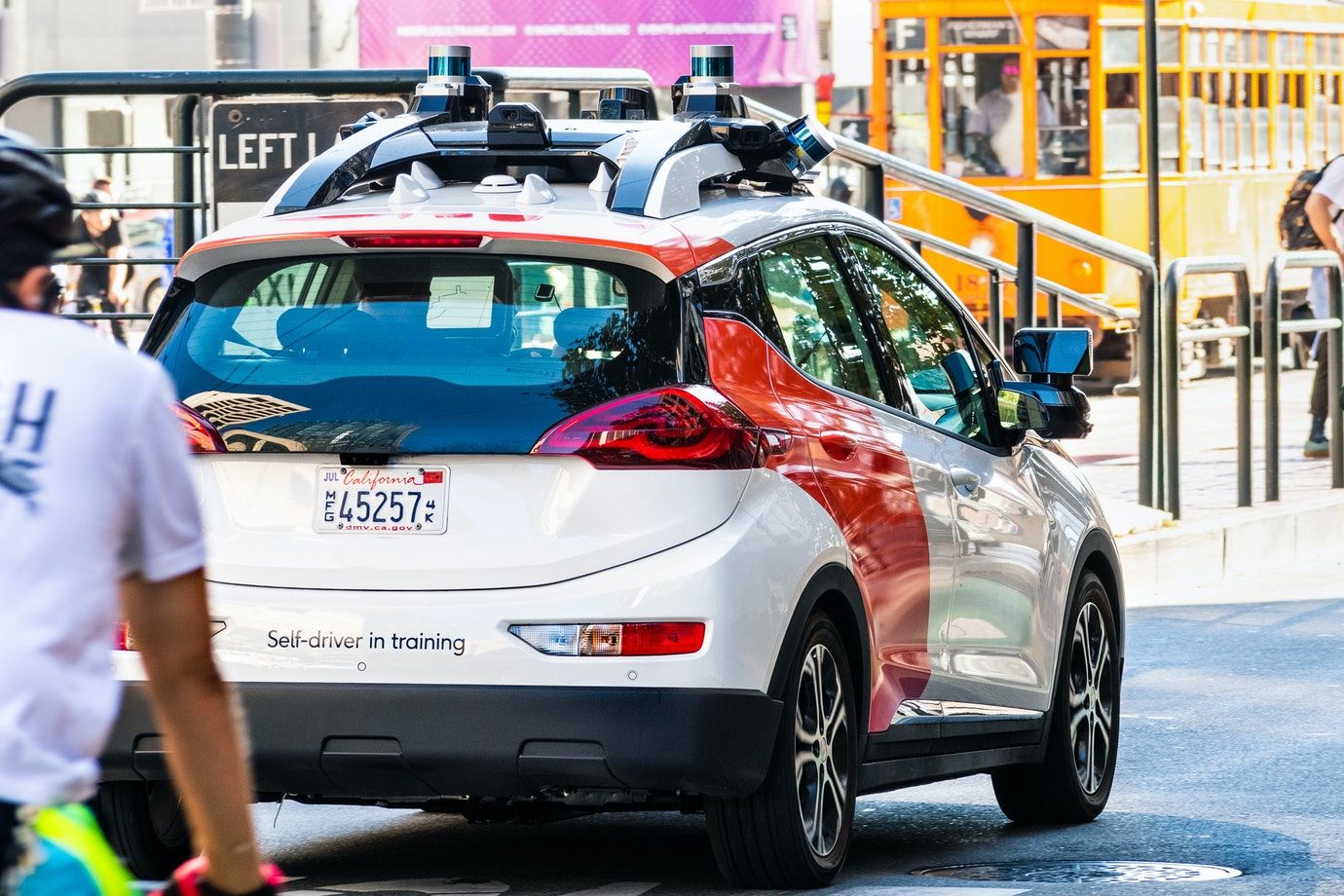 This article is about how self-driving cars and 5G will make roads safer for everyone.
This article is about how self-driving cars and 5G will make roads safer for everyone.
 Unlock business growth with AI, language models, and automation. Learn proven tips for seamless innovation integration into your business strategy.
Unlock business growth with AI, language models, and automation. Learn proven tips for seamless innovation integration into your business strategy.
 The story of Netflix as a company is the story of a business that knows how to innovate, takes chances, and takes risks that even compromise its business model
The story of Netflix as a company is the story of a business that knows how to innovate, takes chances, and takes risks that even compromise its business model
 Artificial Intelligence is going to push development
Artificial Intelligence is going to push development
 If you haven’t been paying attention to digital currencies, now is the time to start.
If you haven’t been paying attention to digital currencies, now is the time to start.
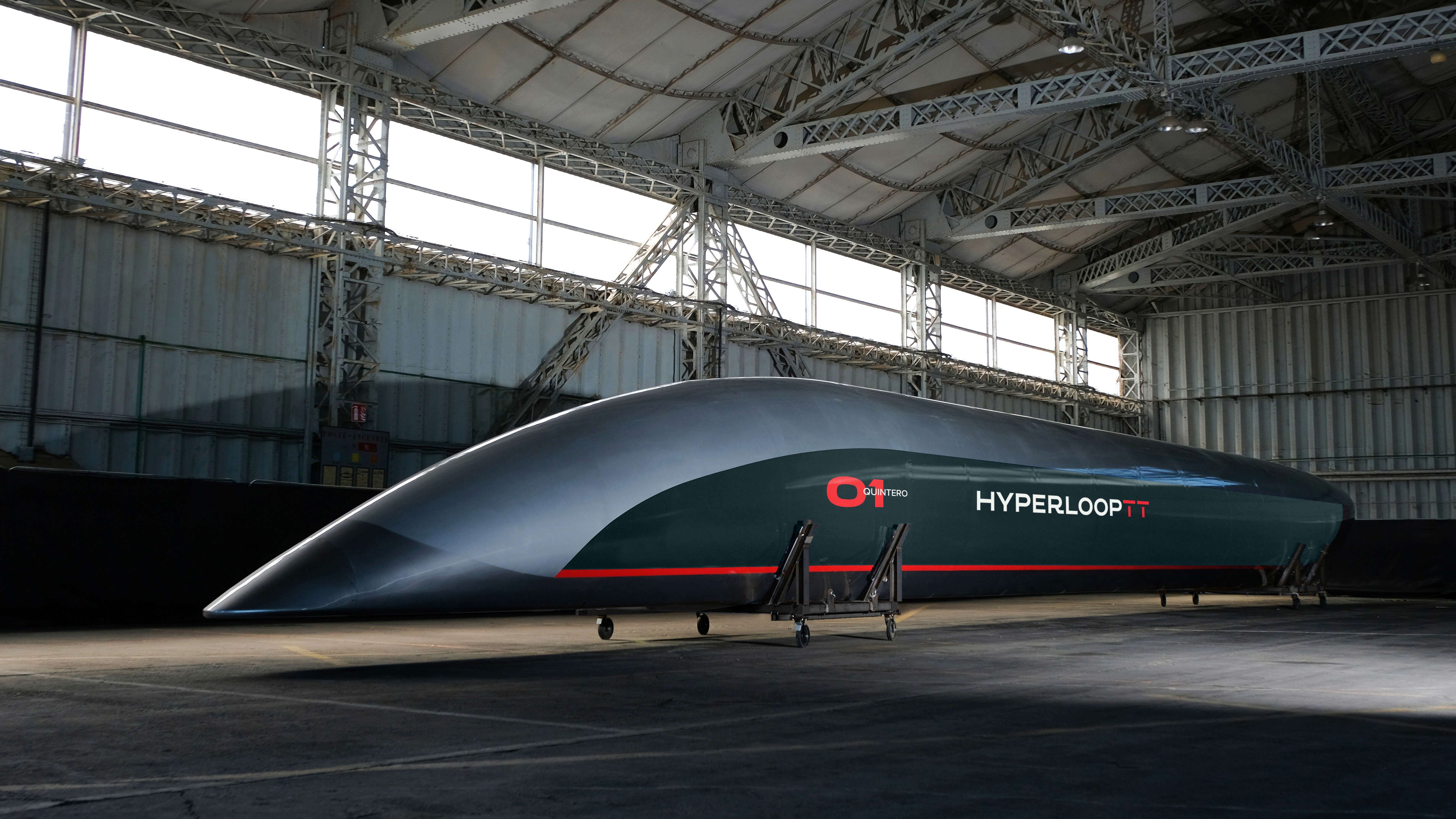 Originally conceived by Elon Musk in 2013, the Hyperloop has been touted as the fastest way to cross the surface of the Earth. Possibly one of the greatest leaps in transportation for generations, the concept promises to slash journey times between cities from several hours to a matter of minutes. On track to revolutionise our world, when can we expect the Hyperloop to become a reality and is it too good to be true?
Originally conceived by Elon Musk in 2013, the Hyperloop has been touted as the fastest way to cross the surface of the Earth. Possibly one of the greatest leaps in transportation for generations, the concept promises to slash journey times between cities from several hours to a matter of minutes. On track to revolutionise our world, when can we expect the Hyperloop to become a reality and is it too good to be true?
 Explore the latest in consumer brain devices for meditation, gaming, and more. Compare top devices of 2024 and discover the perfect BCI for your needs!
Explore the latest in consumer brain devices for meditation, gaming, and more. Compare top devices of 2024 and discover the perfect BCI for your needs!
 The US Defense Advanced Research Projects Agency (DARPA) is looking to improve traumatic memory consolidation by modulating REM sleep for stress and trauma.
The US Defense Advanced Research Projects Agency (DARPA) is looking to improve traumatic memory consolidation by modulating REM sleep for stress and trauma.
 Margin trading magnifies the market’s every move. Leverage intensifies the trading experience and captivates our inner risk-taker. It’s either moon or gloom. We are lured in, naturally focusing on the upside potential. As volumes grow along with increasing crypto-optimism, demand for leveraged exposure is rising.
Margin trading magnifies the market’s every move. Leverage intensifies the trading experience and captivates our inner risk-taker. It’s either moon or gloom. We are lured in, naturally focusing on the upside potential. As volumes grow along with increasing crypto-optimism, demand for leveraged exposure is rising.
 So, first the disclaimer…
So, first the disclaimer…
 Ever since first contender blockchains emerged decade ago, the consensus debate has raged on. The overall concept of proof-of-work (PoW) pre-dates Satoshi Nakamoto’s white paper by around 15 years. However, Satoshi was the trailblazer of using PoW in a decentralized peer-to-peer environment. He was the first to propose it as a means of economic incentive for preventing the double-spend problem.
Ever since first contender blockchains emerged decade ago, the consensus debate has raged on. The overall concept of proof-of-work (PoW) pre-dates Satoshi Nakamoto’s white paper by around 15 years. However, Satoshi was the trailblazer of using PoW in a decentralized peer-to-peer environment. He was the first to propose it as a means of economic incentive for preventing the double-spend problem.
 It is fun to speculate on the price of Bitcoin or whatever your favorite asset is, but the real question for our industry is what value are we delivering to our customers. How are we making their lives better, saving them time or money? If a crypto exchange needs a major bull run in Bitcoin to survive, then they have the wrong business model.
It is fun to speculate on the price of Bitcoin or whatever your favorite asset is, but the real question for our industry is what value are we delivering to our customers. How are we making their lives better, saving them time or money? If a crypto exchange needs a major bull run in Bitcoin to survive, then they have the wrong business model.
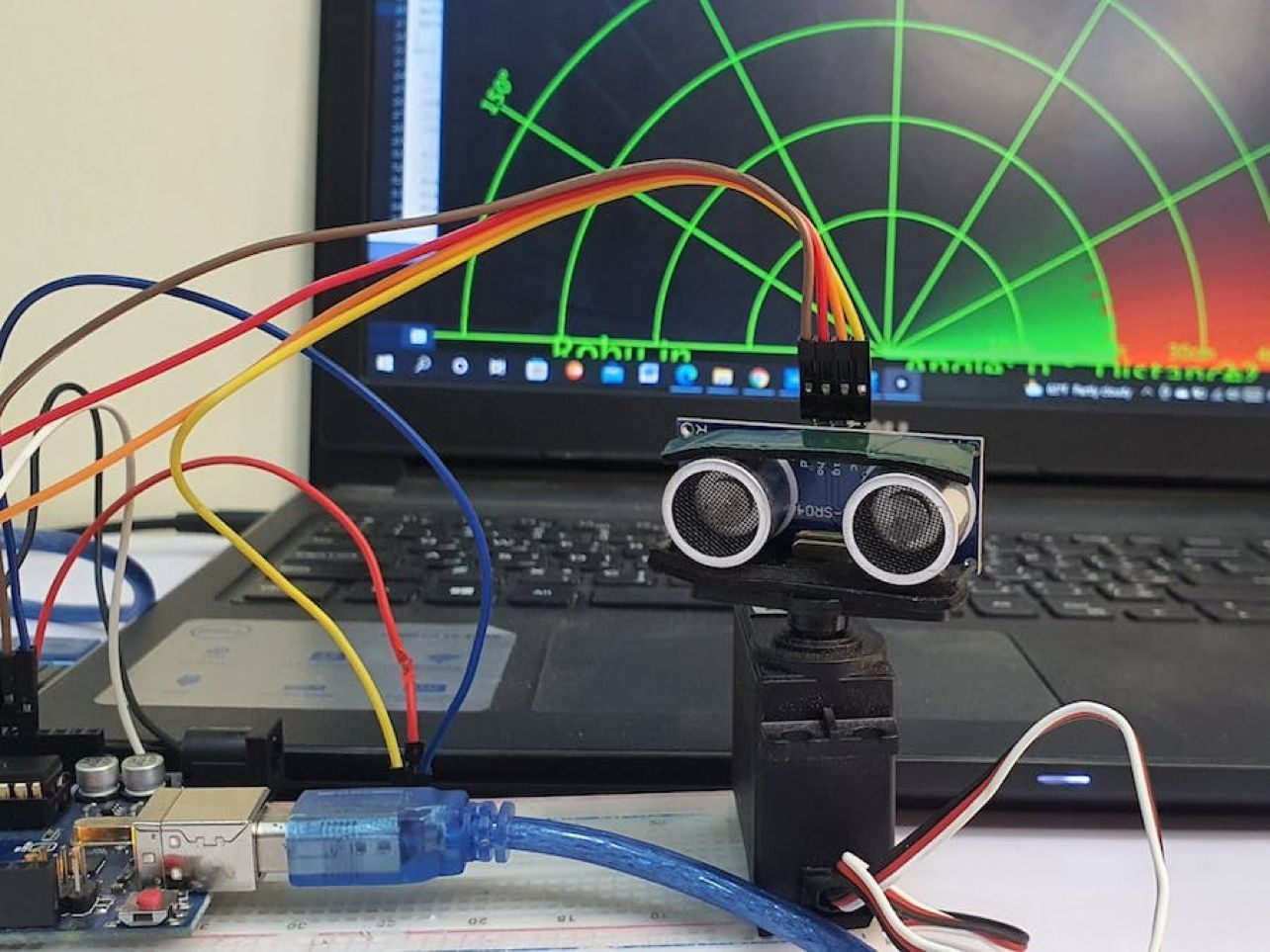 Despite the recent market shocks, blockchain is bet upon by many to bring innovation to finance, tech, and science.
Despite the recent market shocks, blockchain is bet upon by many to bring innovation to finance, tech, and science.
 The corona virus has challenged all aspects of our lives. Healthcare not with standing, one of the biggest challenges has been in trying to keep as much of our lives as possible running as normal. Technology might already have altered the way we work, rest and play for good– but it’s been even more crucial during a period where people are working from home and avoiding large gatherings in the US and the rest of the world. In this post, we’ll look at how tech industry is rising to the corona virus challenge to keep the world moving.
The corona virus has challenged all aspects of our lives. Healthcare not with standing, one of the biggest challenges has been in trying to keep as much of our lives as possible running as normal. Technology might already have altered the way we work, rest and play for good– but it’s been even more crucial during a period where people are working from home and avoiding large gatherings in the US and the rest of the world. In this post, we’ll look at how tech industry is rising to the corona virus challenge to keep the world moving.
 ‘Ethereum killers’ are largely a media narrative - with Solana, Polygon, and Harmony as the only outperformers
‘Ethereum killers’ are largely a media narrative - with Solana, Polygon, and Harmony as the only outperformers
 Working in Sales for the past 5 years means the browser tab I most consistently have open is my email. It's also one of the very few active app notifications I have on my phone.
Working in Sales for the past 5 years means the browser tab I most consistently have open is my email. It's also one of the very few active app notifications I have on my phone.
 Blockchain will transform the internet and the way we use it. From digital freedom to data protection; the reasons are becoming more important every day.
Blockchain will transform the internet and the way we use it. From digital freedom to data protection; the reasons are becoming more important every day.
 Explore NEOM's visionary ambitions, potential, challenges, and global impact, and decide for yourself whether smart cities are the future.
Explore NEOM's visionary ambitions, potential, challenges, and global impact, and decide for yourself whether smart cities are the future.
 How to build a startup without learning to code
How to build a startup without learning to code
 Digital Technology is everywhere and it is redefining how we live, communicate, and work. Most importantly, it accelerates how we innovate.
Digital Technology is everywhere and it is redefining how we live, communicate, and work. Most importantly, it accelerates how we innovate.
 Imagine if any kind of blockchain you are currently looking at is a 2D point of view of a 3D shaped blockchain. If you want to discover more read my story !
Imagine if any kind of blockchain you are currently looking at is a 2D point of view of a 3D shaped blockchain. If you want to discover more read my story !
 Digital Transformation and AI in specific has evolved the business industry drastically. And now it’s time to evolve your recruitment process with AI.
Digital Transformation and AI in specific has evolved the business industry drastically. And now it’s time to evolve your recruitment process with AI.
 Stop chasing last season's gold rush. Discover why prerequisites are a myth, why your network is a side effect of your work, and where new worlds are built
Stop chasing last season's gold rush. Discover why prerequisites are a myth, why your network is a side effect of your work, and where new worlds are built
 I believe that anything we do needs to be grounded in theory, otherwise it’s just a game of numbers—us doing random things and hoping that something will stick.
I believe that anything we do needs to be grounded in theory, otherwise it’s just a game of numbers—us doing random things and hoping that something will stick.
 The digital ‘economy’ has now amassed an estimated value of $11.5 trillion globally, growing two and a half times faster than the global GDP across the past…
The digital ‘economy’ has now amassed an estimated value of $11.5 trillion globally, growing two and a half times faster than the global GDP across the past…
 A Startup Model for Corporate Innovation
A Startup Model for Corporate Innovation
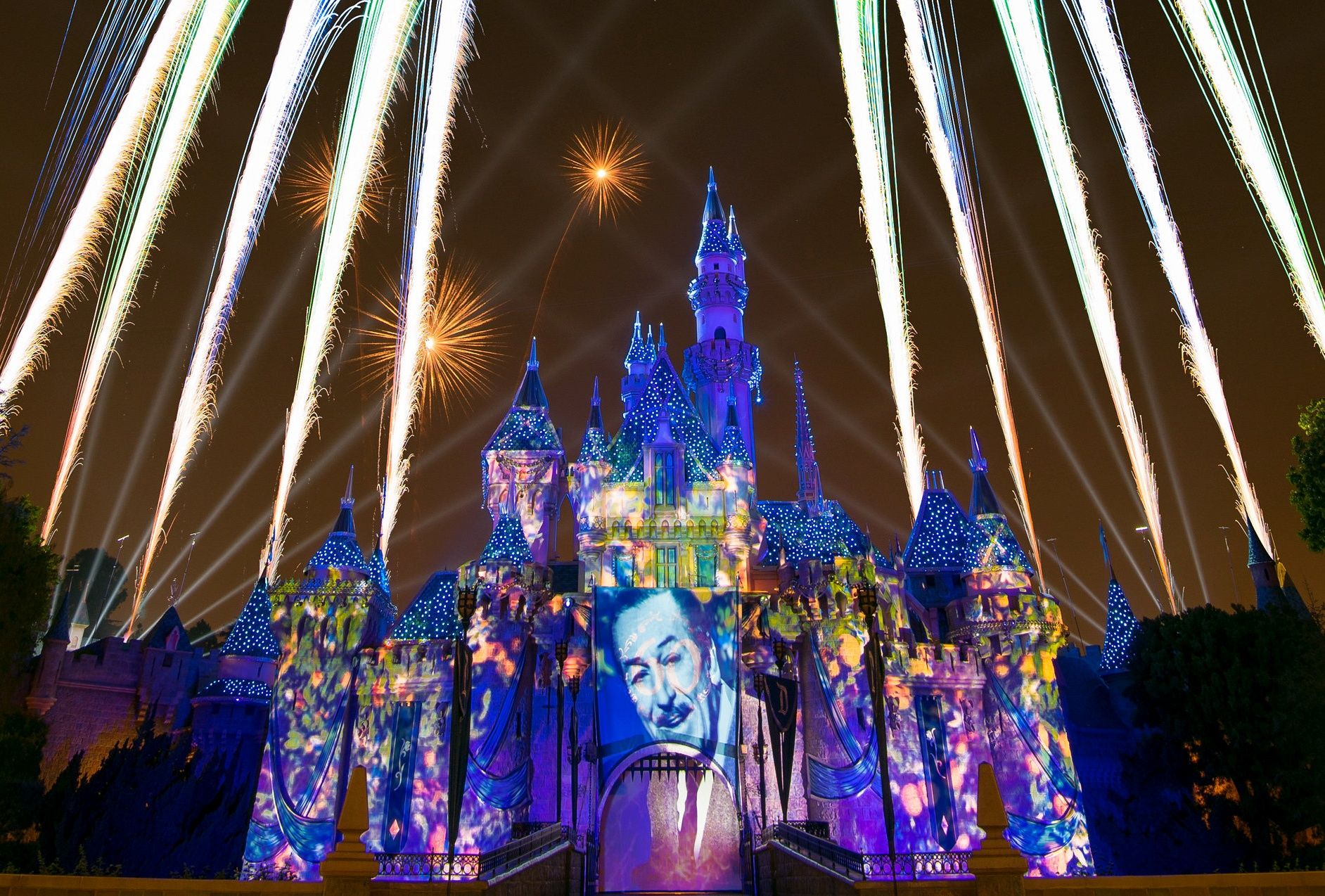 Disney is transforming its theme parks from a physical experience into a theme park Metaverse in a union of in-person and virtual entertainment.
Disney is transforming its theme parks from a physical experience into a theme park Metaverse in a union of in-person and virtual entertainment.
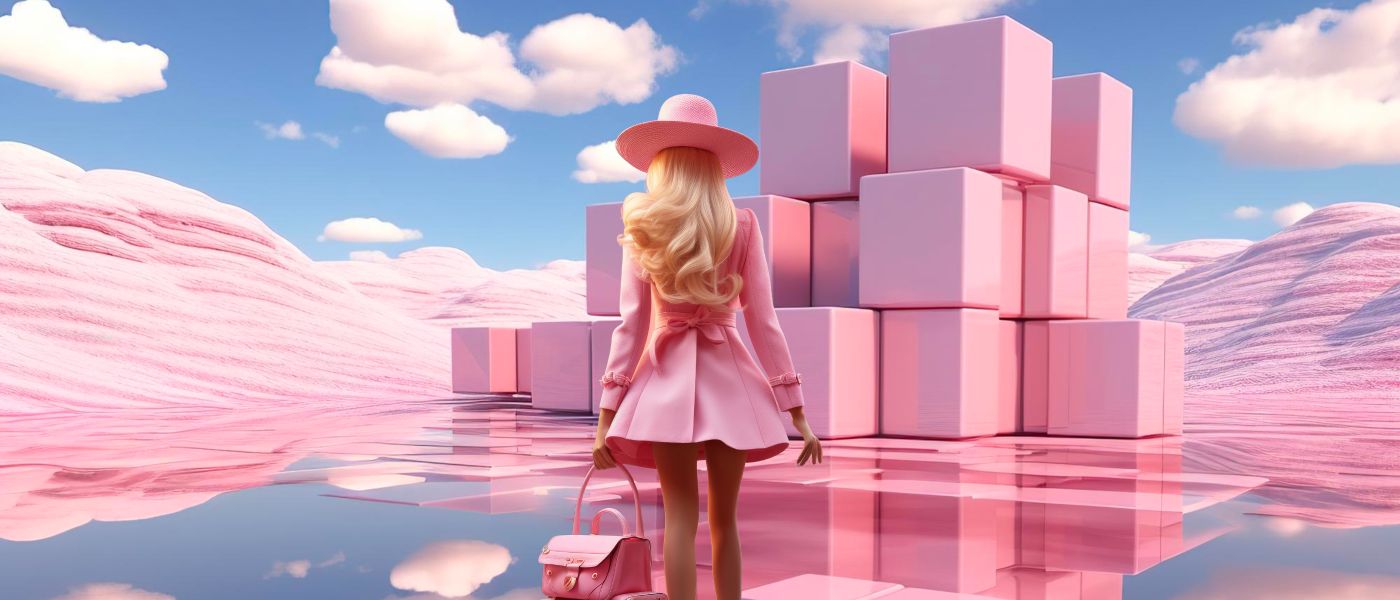 Discover how Barbie's dynamic roles and blockchain's cutting-edge innovations intertwine, emphasizing transformation, inclusivity, and continuous evolution.
Discover how Barbie's dynamic roles and blockchain's cutting-edge innovations intertwine, emphasizing transformation, inclusivity, and continuous evolution.
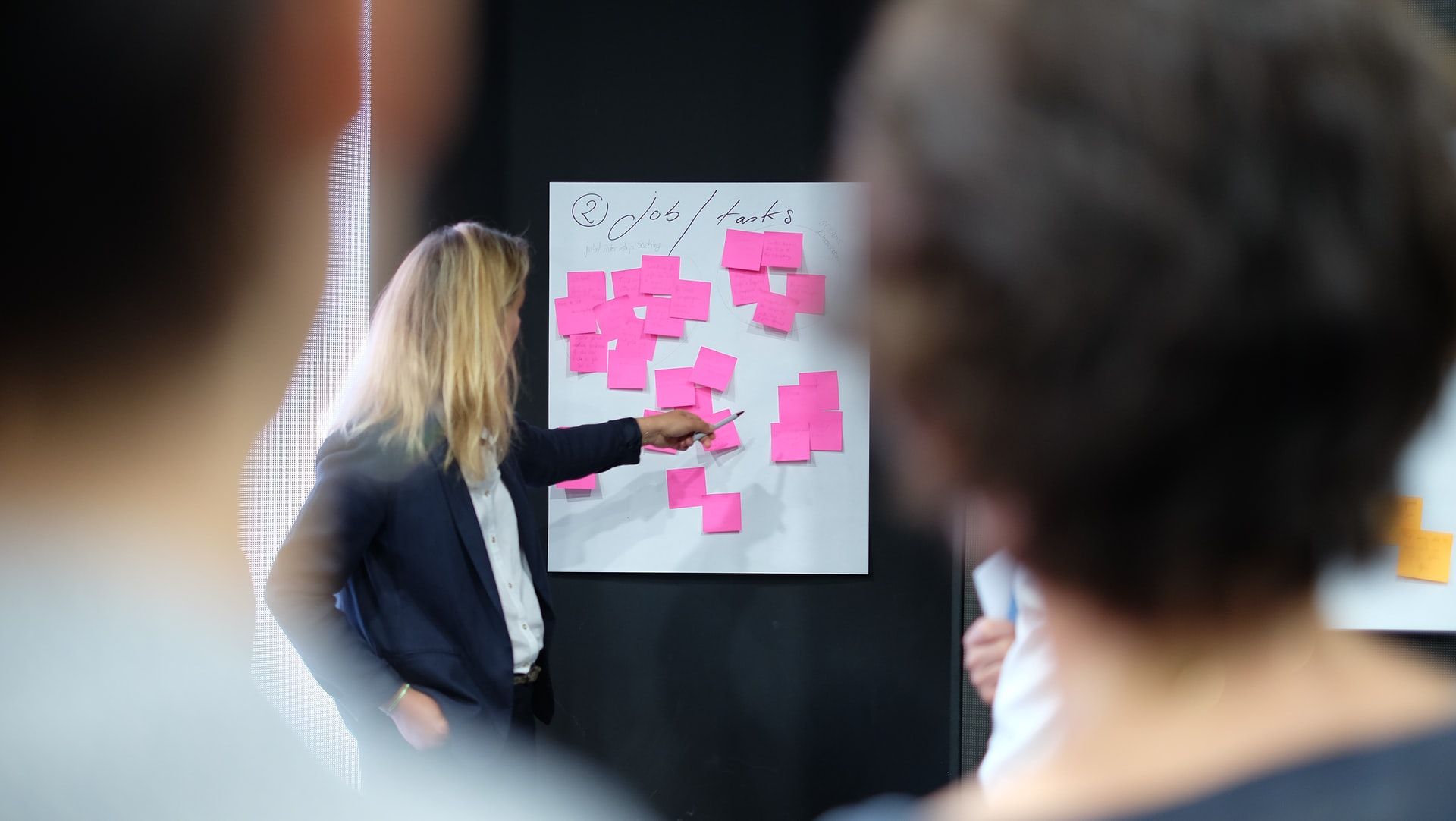 What are the key leadership qualities that inspire teams to build amazing products?
What are the key leadership qualities that inspire teams to build amazing products?
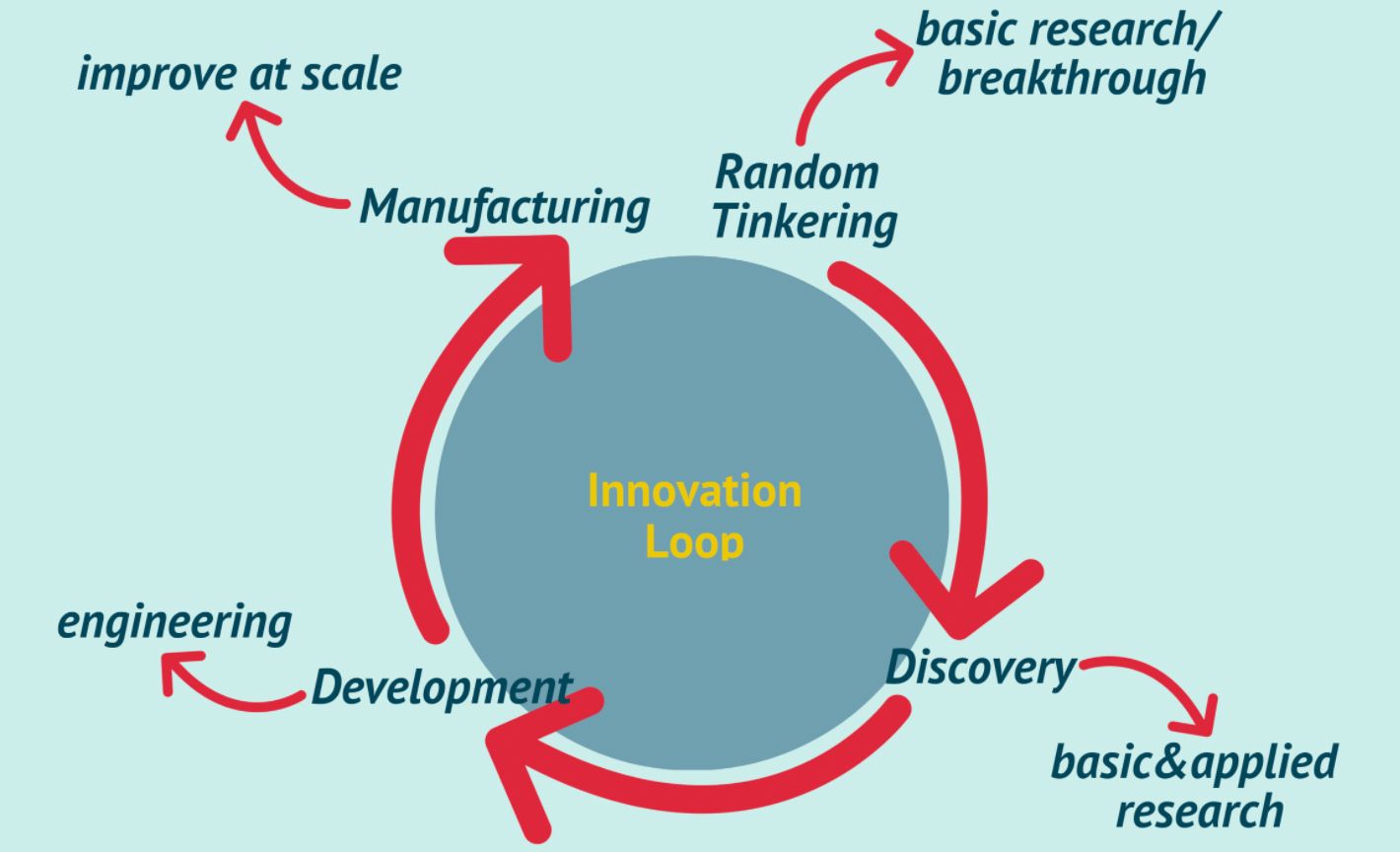 The innovation loop is a methodology/framework derived from the Bell Labs, which produced innovation at scale throughout the 19th century. They learned how to l
The innovation loop is a methodology/framework derived from the Bell Labs, which produced innovation at scale throughout the 19th century. They learned how to l
 Project DNA helps teams execute projects faster, reduce latency, and turn uncertainty into insight, creating a practical framework for innovation success.
Project DNA helps teams execute projects faster, reduce latency, and turn uncertainty into insight, creating a practical framework for innovation success.
 How crypto projects can finally combat legacy social media platforms with the help of data innovation on the blockchain.
How crypto projects can finally combat legacy social media platforms with the help of data innovation on the blockchain.
 Why always looking for a twist or a difference makes all the difference when marketing your products and trying to grow your brand especially at the start.
Why always looking for a twist or a difference makes all the difference when marketing your products and trying to grow your brand especially at the start.
 Discover how visual mind maps, git-like versioning, prompt optimization, and PDF export transform LLM chat apps for efficient, organized research.
Discover how visual mind maps, git-like versioning, prompt optimization, and PDF export transform LLM chat apps for efficient, organized research.
 The days of “dumb” analog devices are at an end. These days, everything has to be “smart” and a part of the Internet-of-Things (IoT).
The days of “dumb” analog devices are at an end. These days, everything has to be “smart” and a part of the Internet-of-Things (IoT).
 Although Hartwell’s statement may initially appear to be drowning in hyperbole, we may one day look back at his words as being precisely accurate.
Although Hartwell’s statement may initially appear to be drowning in hyperbole, we may one day look back at his words as being precisely accurate.
 Clayton Christensen is one of the greatest business minds of our time. His recent passing caused me to reflect on all I learned from him. His life was dedicated to his family, his faith, and the theories he taught. He personified servant leadership.
Clayton Christensen is one of the greatest business minds of our time. His recent passing caused me to reflect on all I learned from him. His life was dedicated to his family, his faith, and the theories he taught. He personified servant leadership.
 According to the World Economic Forum, in 2020 the entire digital universe has reached 44 Zetabytes of Data.
According to the World Economic Forum, in 2020 the entire digital universe has reached 44 Zetabytes of Data.
 Struggling with OSCP exam? These brain-based tips and creative thinking strategies might be the edge you need to break through and think like an innovator.
Struggling with OSCP exam? These brain-based tips and creative thinking strategies might be the edge you need to break through and think like an innovator.
 Former CEO says his company had the opportunity to invest in bitcoin mining but the proposal never came to the board.
Former CEO says his company had the opportunity to invest in bitcoin mining but the proposal never came to the board.
 Unprecedented.
Unprecedented.
 In 2004, I was eager for a challenge. I embarked on an adventure of completing a Computer Science Bachelors degree. Fast forward to today and the adventure continues. I’ve been building products for 12 years for a variety of industries from finance to ecommerce, retail, real estate, hospitality and more.
In 2004, I was eager for a challenge. I embarked on an adventure of completing a Computer Science Bachelors degree. Fast forward to today and the adventure continues. I’ve been building products for 12 years for a variety of industries from finance to ecommerce, retail, real estate, hospitality and more.
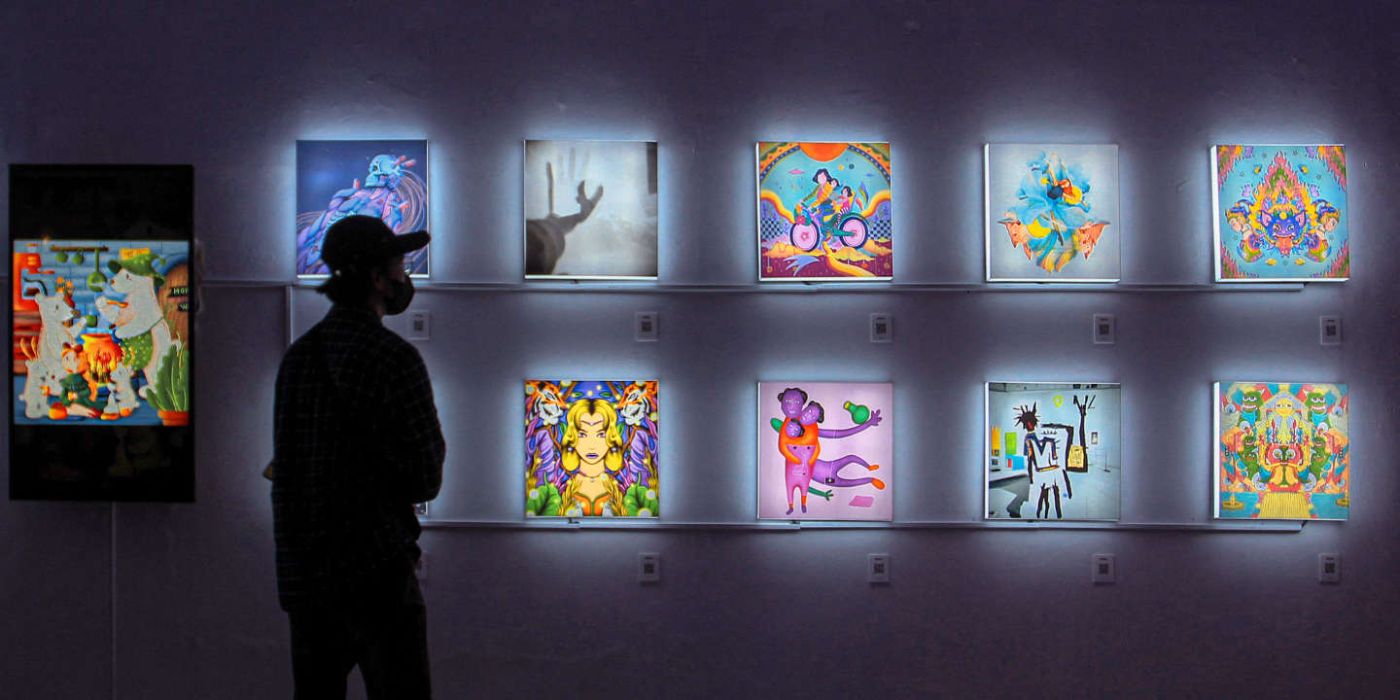 Artist and provocateur Brad Troemel recently stated in an Instagram slideshow post that the present state of NFT art is best defined as "visual dogshit."
Artist and provocateur Brad Troemel recently stated in an Instagram slideshow post that the present state of NFT art is best defined as "visual dogshit."
 Explore the truth about Bitcoin mining's environmental impact, debunking myths and highlighting its positive contributions to energy efficiency and innovation.
Explore the truth about Bitcoin mining's environmental impact, debunking myths and highlighting its positive contributions to energy efficiency and innovation.
 Blockchain technology is disrupting every business industry from finance to music.
Blockchain technology is disrupting every business industry from finance to music.
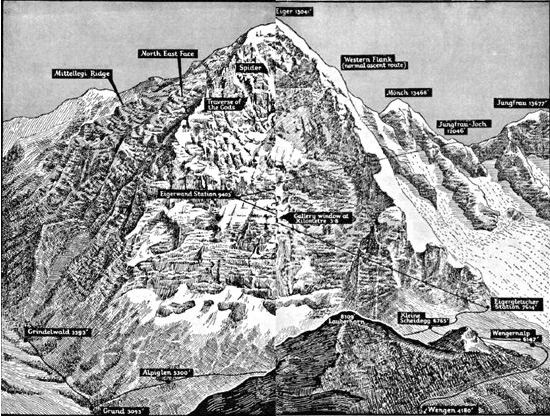 During the 1930s, there was a frenzy for mountain climbing in Europe, driven by some advances in technology, and a modernist zeal. Pre-eminent among these grand challenges, was the lust to climb the North Face of the Eiger, a mile-high sheer face of rock, the largest in the Alps, perpetually in shadow, and shrouded near the top by the dreaded “White Spider” ice field.
During the 1930s, there was a frenzy for mountain climbing in Europe, driven by some advances in technology, and a modernist zeal. Pre-eminent among these grand challenges, was the lust to climb the North Face of the Eiger, a mile-high sheer face of rock, the largest in the Alps, perpetually in shadow, and shrouded near the top by the dreaded “White Spider” ice field.
 Cloud computing has been very boring for a long time. Everything is inside the data center and you don’t even see the systems that you are working on.
Cloud computing has been very boring for a long time. Everything is inside the data center and you don’t even see the systems that you are working on.
 The question of “what makes the best smart contract platform” really depends on who’s asking it as there are many variables which will taken into account or not depending on who is asking the question. An investor will usually only consider the value of the underlying token, which isn’t necessarily an indicator of “the best.” As we know by now, token prices can rise and fall in minutes off the back of rumors and speculation. Just look at the value of Tron each time Justin Sun ends up in the press.
The question of “what makes the best smart contract platform” really depends on who’s asking it as there are many variables which will taken into account or not depending on who is asking the question. An investor will usually only consider the value of the underlying token, which isn’t necessarily an indicator of “the best.” As we know by now, token prices can rise and fall in minutes off the back of rumors and speculation. Just look at the value of Tron each time Justin Sun ends up in the press.
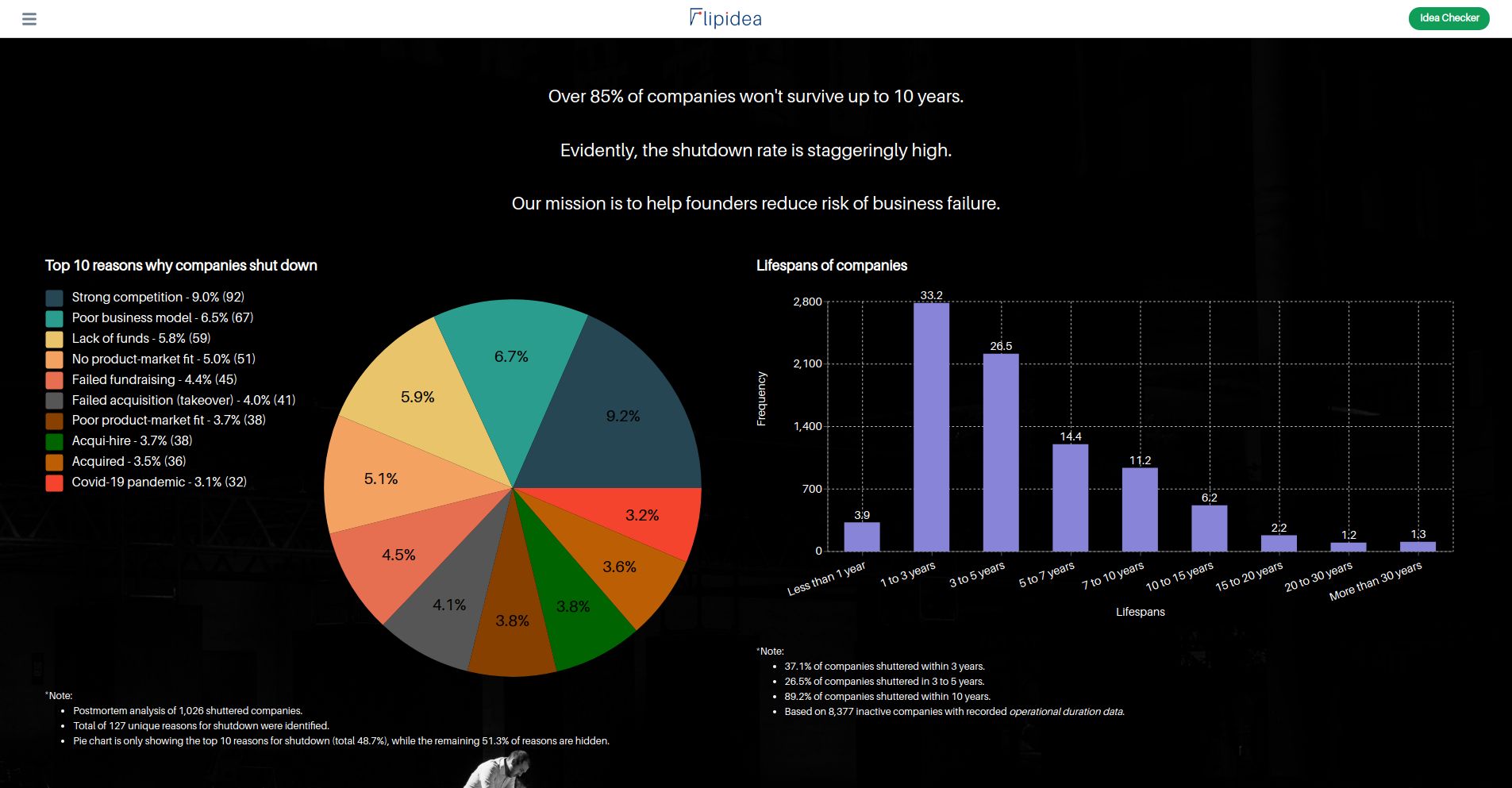 Especially when over 85% of startups won't survive up to 10 years.
Especially when over 85% of startups won't survive up to 10 years.
 how to design a winning new business / venture / product / service, rather than merely ticking off tasks on a to-do list.
how to design a winning new business / venture / product / service, rather than merely ticking off tasks on a to-do list.
 The article explores different cases of integrating "disruptive people" with software and creative teams and how it impacts search for innovation and business continuity.
The article explores different cases of integrating "disruptive people" with software and creative teams and how it impacts search for innovation and business continuity.
 Lithium-ion batteries are the standard for electronics, electric vehicles and are also growing in popularity for military and aerospace applications. A prototype Li-ion battery was developed by Akira Yoshino in 1985, then a commercial Li-ion battery was developed by a Sony and Asahi Kasei team in 1991. Since then, there haven't really been any breakthroughs in rechargeable batteries, however, because Lithium is finite and not renewable, we will need to find different methods of making these batteries that almost all our electronic devices rely on.
Lithium-ion batteries are the standard for electronics, electric vehicles and are also growing in popularity for military and aerospace applications. A prototype Li-ion battery was developed by Akira Yoshino in 1985, then a commercial Li-ion battery was developed by a Sony and Asahi Kasei team in 1991. Since then, there haven't really been any breakthroughs in rechargeable batteries, however, because Lithium is finite and not renewable, we will need to find different methods of making these batteries that almost all our electronic devices rely on.
 A few thoughts from my experience as reader and writer using Medium’s new clap feature the past few weeks:
A few thoughts from my experience as reader and writer using Medium’s new clap feature the past few weeks:
 Photo by Uriel Soberanes on Unsplash
Photo by Uriel Soberanes on Unsplash
 Do It Before It's 'Cool'
Do It Before It's 'Cool'
 Learn everything you need to know about Innovation via these 237 free HackerNoon stories.
Learn everything you need to know about Innovation via these 237 free HackerNoon stories.
 All we talk about is AI. Even your parents talk about it. How is it that we have so quickly accepted an AI-infused future as a fait accompli?
All we talk about is AI. Even your parents talk about it. How is it that we have so quickly accepted an AI-infused future as a fait accompli?
 Software development is not dying it is evolving.
Software development is not dying it is evolving.
 Every tech gets its updation then why not web it also gets its update, there may be a delay for web3 but the wait is worth it.
Every tech gets its updation then why not web it also gets its update, there may be a delay for web3 but the wait is worth it.
 Dating apps stagnate, culture collapses.
Explore how Al concierge could spark a new era of safe, authentic, and meaningful relationships.
Dating apps stagnate, culture collapses.
Explore how Al concierge could spark a new era of safe, authentic, and meaningful relationships.
 First principles thinking drives complex problem solving and workplace innovation through reverse engineering. Companies that employ first principles thinking are one step ahead as they plan and build for the future.
First principles thinking drives complex problem solving and workplace innovation through reverse engineering. Companies that employ first principles thinking are one step ahead as they plan and build for the future.
 Remember great technological advances that changed the lives of thousands of people and the new bets of the future through information technology.
Remember great technological advances that changed the lives of thousands of people and the new bets of the future through information technology.
 Recently, while exploring Google Gemini's image generation for my personal project, I noticed something unsettling.
Recently, while exploring Google Gemini's image generation for my personal project, I noticed something unsettling.
 Connor Crook, CEO of Diamondback Toolbelts, does give the impression that the destiny of your business is squarely in the hands of fate. However …
Connor Crook, CEO of Diamondback Toolbelts, does give the impression that the destiny of your business is squarely in the hands of fate. However …
 Companies like Alphabet have consistently sunk millions into their innovation departments, only to be routinely outpaced by smaller operations.
Companies like Alphabet have consistently sunk millions into their innovation departments, only to be routinely outpaced by smaller operations.
 How Universal Basic Income (UBI) provides food on the table and bills to be paid
How Universal Basic Income (UBI) provides food on the table and bills to be paid
 There’s a formula that unlocks the secret sauce for creating value.
There’s a formula that unlocks the secret sauce for creating value.
 “You have to debug your business model.”
“You have to debug your business model.”
 For businesses to maximize the benefits of innovation, they must consider safety and compliance to avoid a tragedy similar to that of OceanGate.
For businesses to maximize the benefits of innovation, they must consider safety and compliance to avoid a tragedy similar to that of OceanGate.
 Tech innovation has become the driving force of our economy, having an immense contribution to economic growth.
Tech innovation has become the driving force of our economy, having an immense contribution to economic growth.
 Interview with Joey Poareo, co-founder at InuYasha, a decentralized incubator accelerator fund that uses uses DAO governance to speed up crypto projects.
Interview with Joey Poareo, co-founder at InuYasha, a decentralized incubator accelerator fund that uses uses DAO governance to speed up crypto projects.
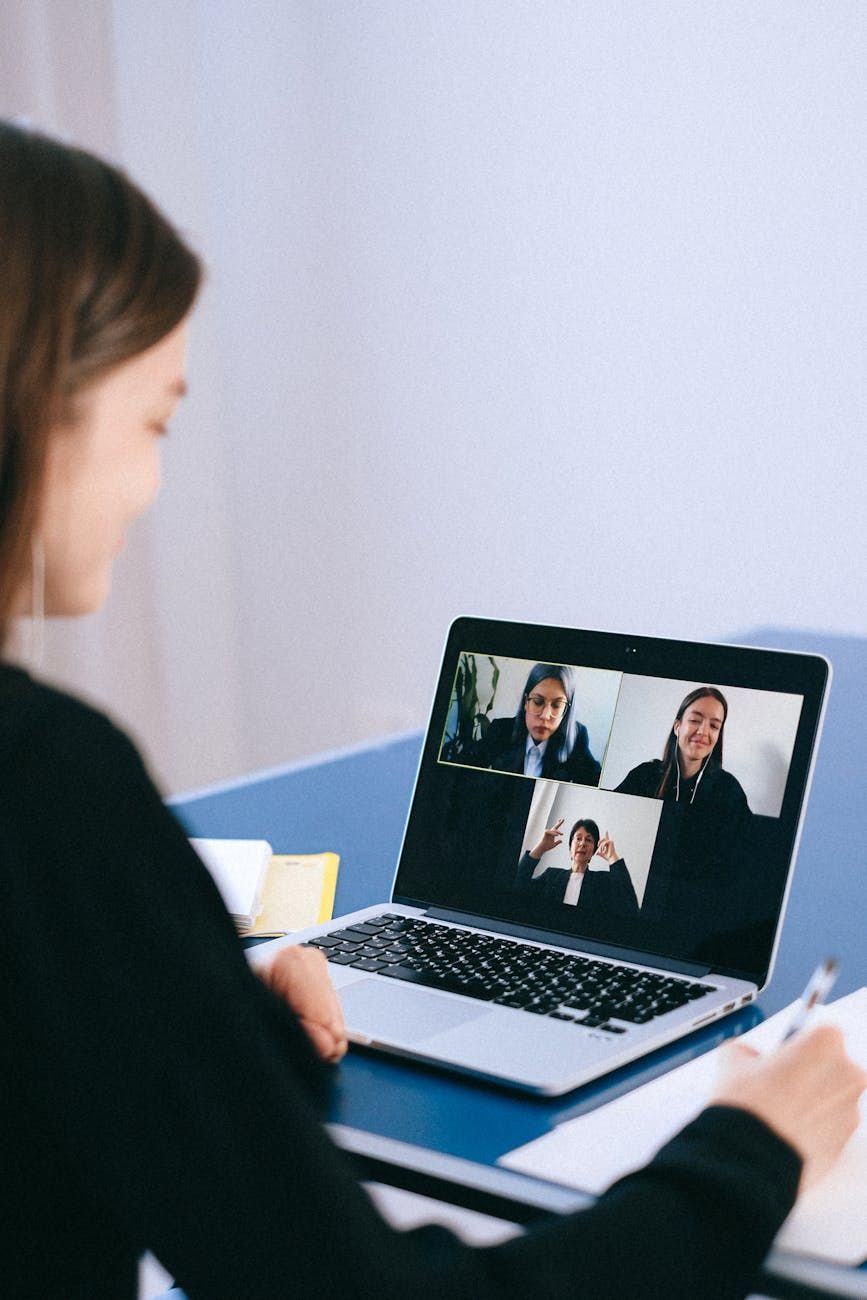 Video conferencing tools like Zoom, Skype, or WebEx along with Digital tools like Microsoft Teams and Slack are deeply embedded into the modern ways of working — even more during the lock-down era. And their role will only get more important.
Video conferencing tools like Zoom, Skype, or WebEx along with Digital tools like Microsoft Teams and Slack are deeply embedded into the modern ways of working — even more during the lock-down era. And their role will only get more important.
 A few days ago I somehow ended up on Google's web.dev platform, which I'm assuming is rather new. There is of course the possibility that I have been–or still am–living under a rock, when it comes to new web technologies.
A few days ago I somehow ended up on Google's web.dev platform, which I'm assuming is rather new. There is of course the possibility that I have been–or still am–living under a rock, when it comes to new web technologies.
 “In these unprecedented times…” People build unprecedented products, and contribute to the internet in unprecedented ways. Go on, make a fellow human’s day and nominate the best YOUR best of 2020’s tech industry for a 2020 #Noonie, the tech industry’s most independent
and community-driven awards: NOONIES.TECH. One such impressive human is Tian Zhao from Canada: 2020 Noonie nominee in Future Heroes and Technology categories.
“In these unprecedented times…” People build unprecedented products, and contribute to the internet in unprecedented ways. Go on, make a fellow human’s day and nominate the best YOUR best of 2020’s tech industry for a 2020 #Noonie, the tech industry’s most independent
and community-driven awards: NOONIES.TECH. One such impressive human is Tian Zhao from Canada: 2020 Noonie nominee in Future Heroes and Technology categories.
 Explore the evolution of process mining since 2011, focusing on advancements by industry leaders like Celonis and UiPath.
Explore the evolution of process mining since 2011, focusing on advancements by industry leaders like Celonis and UiPath.
 The origins of progress and the stagnation of the West
The origins of progress and the stagnation of the West
 As a Software-as-a-Service business, building awareness is imperative for your endeavour. Content marketing has established itself as one of the most powerful tools a SaaS company can use to attract, educate and convert customers.
As a Software-as-a-Service business, building awareness is imperative for your endeavour. Content marketing has established itself as one of the most powerful tools a SaaS company can use to attract, educate and convert customers.
 Explore the transformative landscape of enterprise data intelligence, merging big data, machine learning, and more for streamlined decision-making.
Explore the transformative landscape of enterprise data intelligence, merging big data, machine learning, and more for streamlined decision-making.
 Discover why full-stack, outcome-driven AI companies will dominate 2026. Learn the post-AI growth blueprint, multi-agent systems, and outcome-based strategies.
Discover why full-stack, outcome-driven AI companies will dominate 2026. Learn the post-AI growth blueprint, multi-agent systems, and outcome-based strategies.
 Here's the latest on the growing integration of blockchain technology into the mainstream, showing cryptocurrencies' potential to become a new asset class..
Here's the latest on the growing integration of blockchain technology into the mainstream, showing cryptocurrencies' potential to become a new asset class..
 Growing up in South Africa, the team at Wildcards witnessed first-hand the impact of poaching, pollution, poisoning and other forms of destructive human behavior leading to the extinction and endangerment of hundreds of animal species.
Growing up in South Africa, the team at Wildcards witnessed first-hand the impact of poaching, pollution, poisoning and other forms of destructive human behavior leading to the extinction and endangerment of hundreds of animal species.
 AI has been making headlines quite frequently nowadays. There is a hype and new leap in technology, so let's evaluate it to see what can we expect in 2023.
AI has been making headlines quite frequently nowadays. There is a hype and new leap in technology, so let's evaluate it to see what can we expect in 2023.
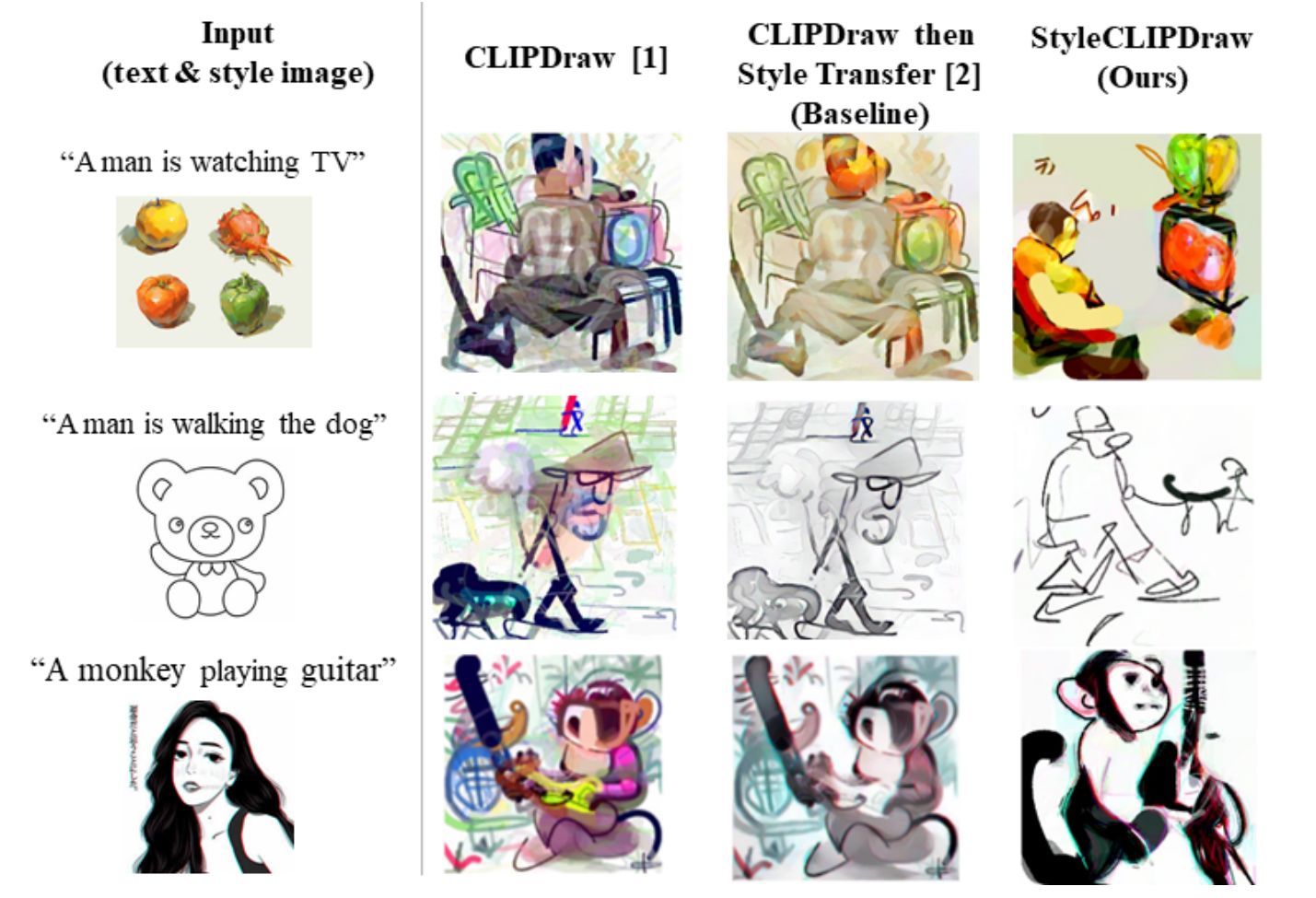 Have you ever dreamed of taking the style of a picture, like this cool TikTok drawing style on the left, and applying it to a new picture of your choice? Well, I did, and it has never been easier to do. In fact, you can even achieve that from only text and can try it right now with this new method and their Google Colab notebook available for everyone (see references).
Have you ever dreamed of taking the style of a picture, like this cool TikTok drawing style on the left, and applying it to a new picture of your choice? Well, I did, and it has never been easier to do. In fact, you can even achieve that from only text and can try it right now with this new method and their Google Colab notebook available for everyone (see references).
 Shiny hardware will get the applause at CES 2026. But for these two categories, I'm betting software's what will bring in cold, hard cash.
Shiny hardware will get the applause at CES 2026. But for these two categories, I'm betting software's what will bring in cold, hard cash.
 Unlock innovative product design strategies in Designing in the Dark by Angelina Severino. Embrace creativity from diverse sources for groundbreaking solutions
Unlock innovative product design strategies in Designing in the Dark by Angelina Severino. Embrace creativity from diverse sources for groundbreaking solutions
 This article describes the ‘idea model’ and provides various examples of real ideas — explains how to write effective executive summaries for your concepts.
This article describes the ‘idea model’ and provides various examples of real ideas — explains how to write effective executive summaries for your concepts.
 Companies are pressed to innovate their applications and software to meet changing consumer and market demands, but it is not happening as fast as it should.
Companies are pressed to innovate their applications and software to meet changing consumer and market demands, but it is not happening as fast as it should.
 Data science is a rapidly developing sector of study. Its main goal is to translate vast amounts of records into valuable business insights. Implementing data science-based tools into your company can be highly beneficial. AI software is more efficient and accurate than humans have ever been.
Data science is a rapidly developing sector of study. Its main goal is to translate vast amounts of records into valuable business insights. Implementing data science-based tools into your company can be highly beneficial. AI software is more efficient and accurate than humans have ever been.
 Raj Jaiswal is the founder and CEO of Neyon, an e-commerce startup. He talks about how he blended tech acumen with business savvy.
Raj Jaiswal is the founder and CEO of Neyon, an e-commerce startup. He talks about how he blended tech acumen with business savvy.
 The past few months of COVID-19 has all but certainly put the nail in the coffin for WeWork. In addition to the stress placed on the U.S.
The past few months of COVID-19 has all but certainly put the nail in the coffin for WeWork. In addition to the stress placed on the U.S.
 The DeFi ecosystem is growing fast and needs more efficient solutions. Perpetual Constant Maturity Options is a step forward towards the Future of DeFi.
The DeFi ecosystem is growing fast and needs more efficient solutions. Perpetual Constant Maturity Options is a step forward towards the Future of DeFi.
 Mirav Vyas has advised a multitude of product and innovation teams across the UK, US and Africa over the last 8+ years.
Mirav Vyas has advised a multitude of product and innovation teams across the UK, US and Africa over the last 8+ years.
 A clearly articulated purpose is only effective when accompanied with an authentic process.
A clearly articulated purpose is only effective when accompanied with an authentic process.
 An interview with Naftali Dratman, CEO of Israel Innovation.
An interview with Naftali Dratman, CEO of Israel Innovation.
 Many innovations need time to catch on because of our resistance and scepticism to a novel way to accomplish things.
Many innovations need time to catch on because of our resistance and scepticism to a novel way to accomplish things.
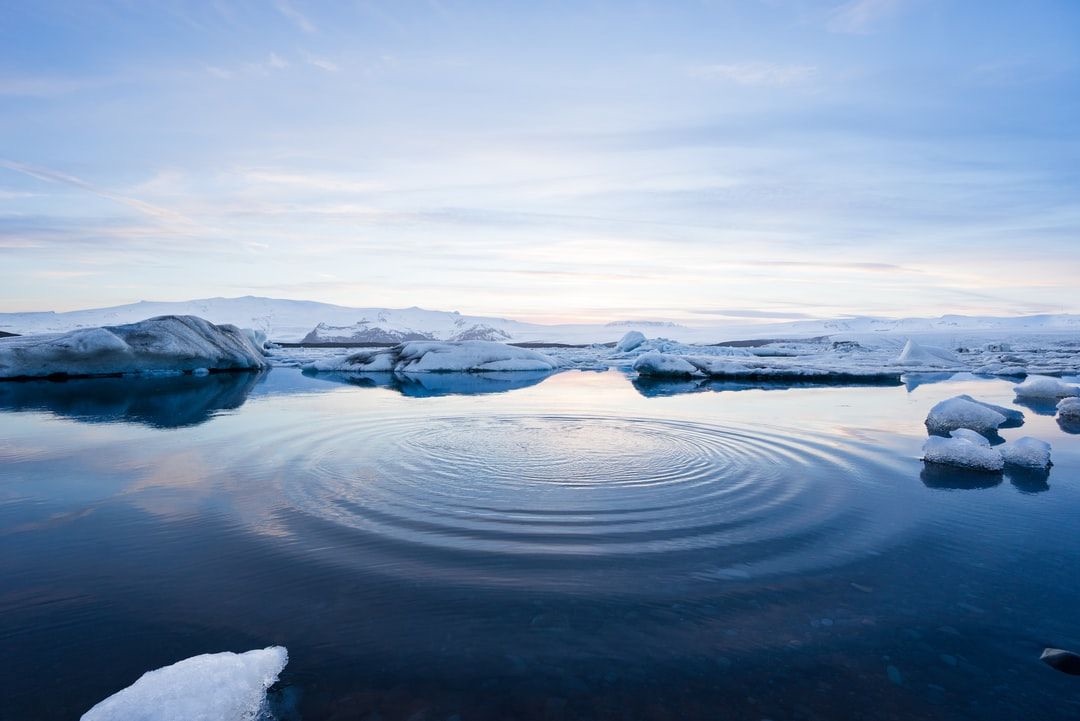 A Story About Socialist Yugoslavia's DIY Computer Revolution in the 1980s.
A Story About Socialist Yugoslavia's DIY Computer Revolution in the 1980s.
 Sarat Chandra Routhu shares his journey in IT services, product development, and privacy compliance, driving innovation and building scalable solutions.
Sarat Chandra Routhu shares his journey in IT services, product development, and privacy compliance, driving innovation and building scalable solutions.
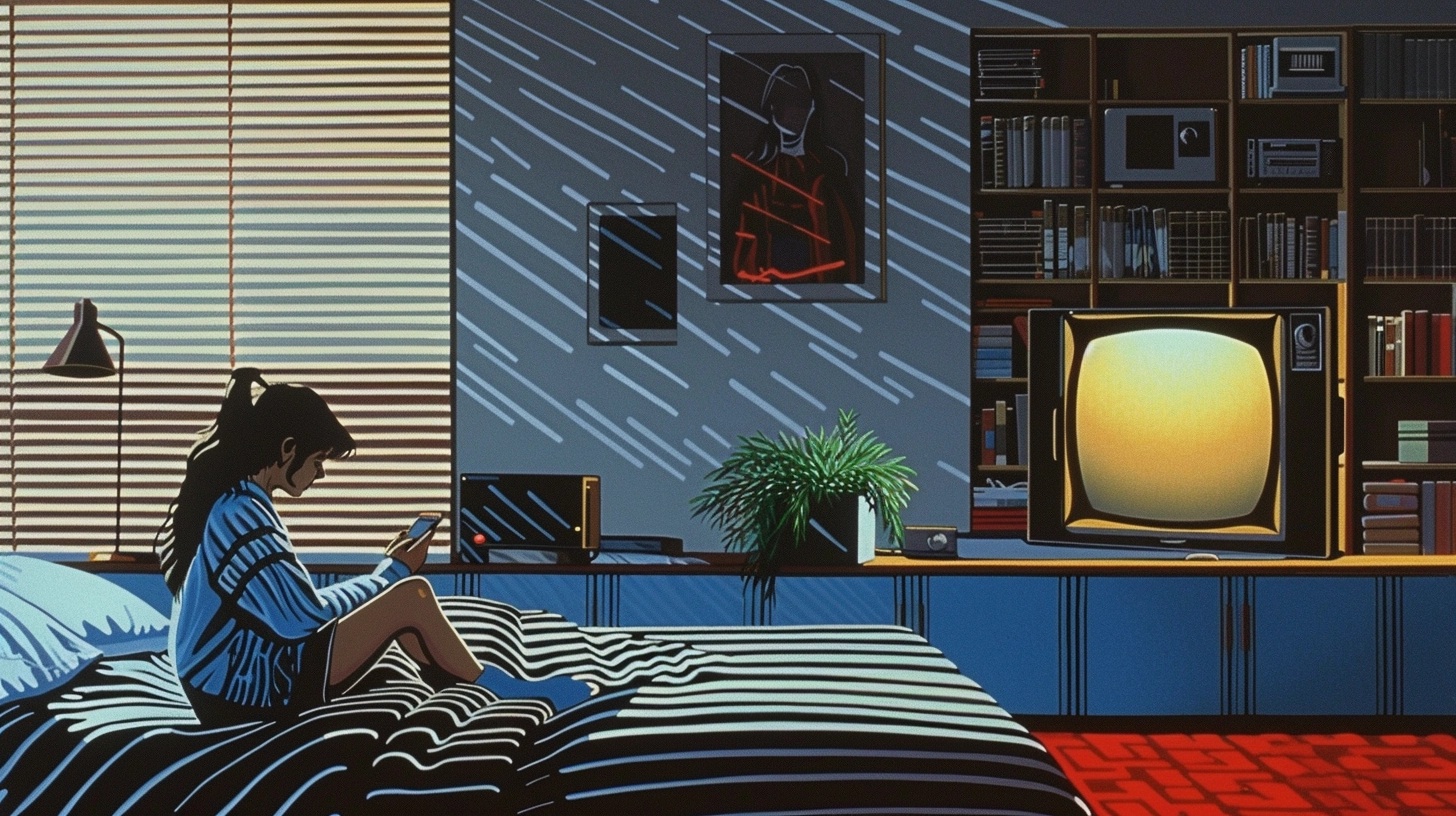 An informative article about unhealthy ways that exacerbate loneliness and innovative tech companies that are making efforts to fight the loneliness epidemic.
An informative article about unhealthy ways that exacerbate loneliness and innovative tech companies that are making efforts to fight the loneliness epidemic.
 These dreams go on when I close my eyes
Every second of the night I live another life
These dreams that sleep when it’s cold outside
Every moment I’m awake the further I’m away
These dreams go on when I close my eyes
Every second of the night I live another life
These dreams that sleep when it’s cold outside
Every moment I’m awake the further I’m away
 Piracy motivates software companies to spend resources on research.
Piracy motivates software companies to spend resources on research.
 Have you ever tasted a veggie burger? Are you a vegan? You have a vegan friend, should you turn vegan? Did you ever thought about stop eating meat? Are you aware of animal slaughtering?
Have you ever tasted a veggie burger? Are you a vegan? You have a vegan friend, should you turn vegan? Did you ever thought about stop eating meat? Are you aware of animal slaughtering?
 Discover how AI-native browsers like Perplexity Comet are transforming web automation with intelligent agents and reshaping modern business workflows.
Discover how AI-native browsers like Perplexity Comet are transforming web automation with intelligent agents and reshaping modern business workflows.
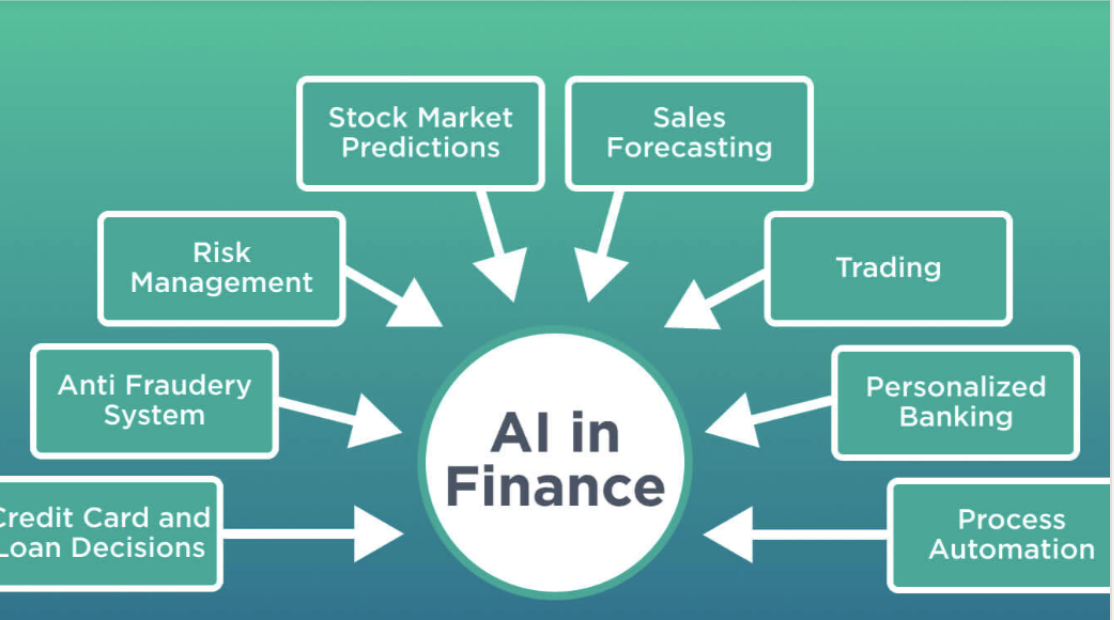 AI isn’t just disrupting financial services—it’s creating a cognitive divide that few are prepared for.
AI isn’t just disrupting financial services—it’s creating a cognitive divide that few are prepared for.
 Restaurants have been hit hard by COVID-19. With massive closures and layoffs around the world, it's been a tough road for hospitality these past few weeks. It can be extremely overwhelming to look at the big picture, but luckily, there are restaurant tech innovators stepping up to the plate.
Restaurants have been hit hard by COVID-19. With massive closures and layoffs around the world, it's been a tough road for hospitality these past few weeks. It can be extremely overwhelming to look at the big picture, but luckily, there are restaurant tech innovators stepping up to the plate.
 “What are the greatest inventions of the past 1,000 years?”
“What are the greatest inventions of the past 1,000 years?”
 10/19/2022: Top 5 stories on the Hackernoon homepage!
10/19/2022: Top 5 stories on the Hackernoon homepage!
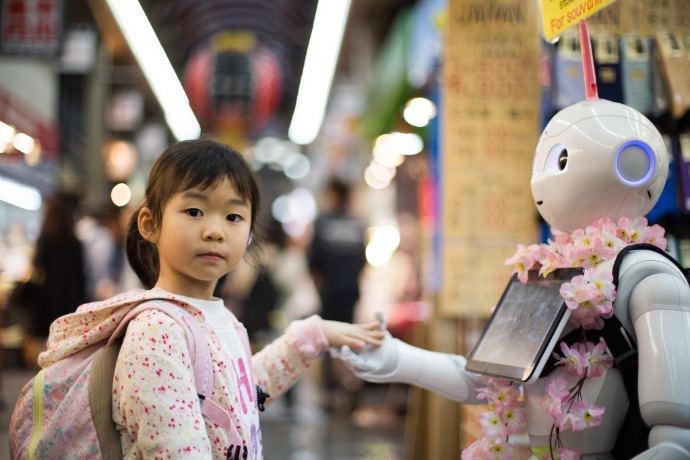 Is AI always responsible for its answers and guidance? Is its undeniable brilliance balanced with a sense of accountability for its actions? Let's find out.
Is AI always responsible for its answers and guidance? Is its undeniable brilliance balanced with a sense of accountability for its actions? Let's find out.
 Working from home — WFH in short — is one of the most debated topics these days. Recent events have compelled many organizations to close their offices and force their employees to work from home. All aspects of the working experience are now being done remotely.
Working from home — WFH in short — is one of the most debated topics these days. Recent events have compelled many organizations to close their offices and force their employees to work from home. All aspects of the working experience are now being done remotely.
 Digital advances are here to aid the Oil and Gas companies in transcending the limitations of traditional techniques.
Digital advances are here to aid the Oil and Gas companies in transcending the limitations of traditional techniques.
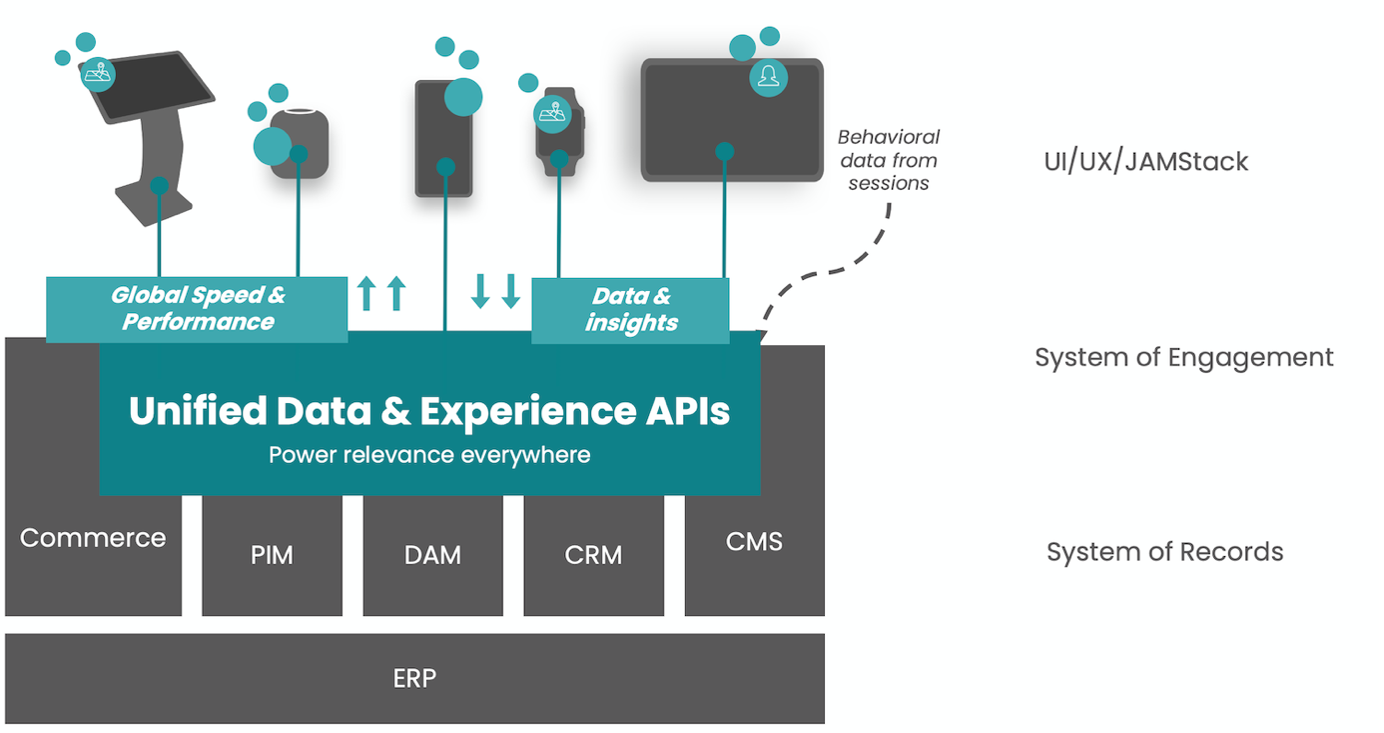 What is it that's holding back retailers from riding on top of the digital disruption wave? The time it takes to integrate software is one of those things.
What is it that's holding back retailers from riding on top of the digital disruption wave? The time it takes to integrate software is one of those things.
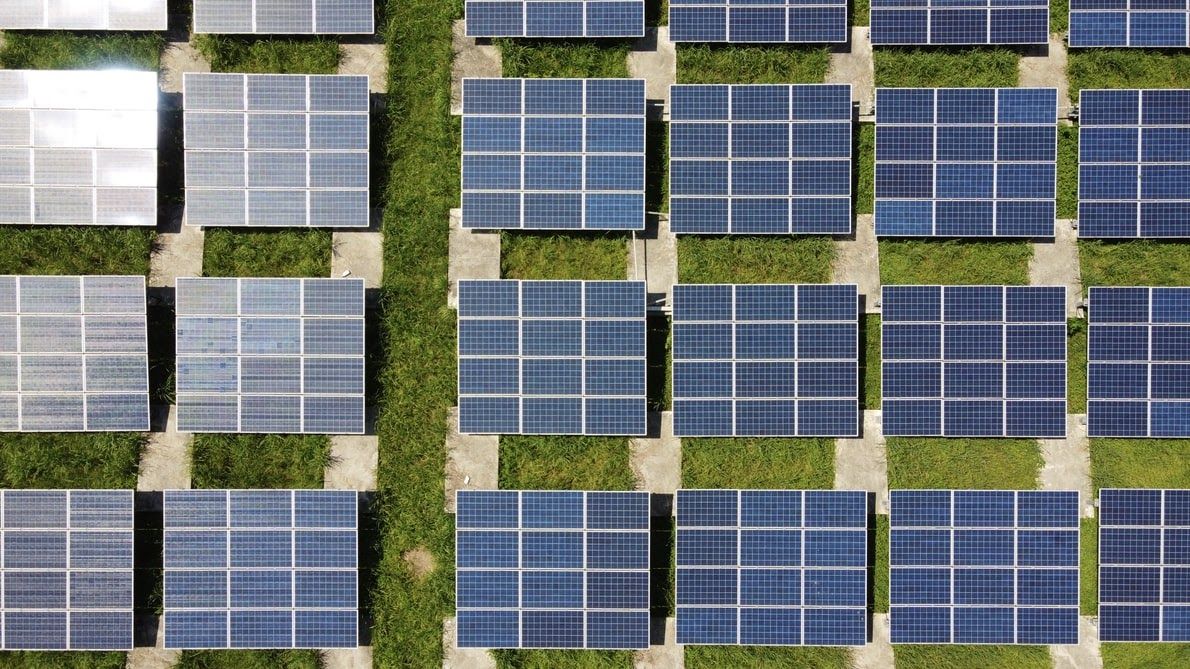 From electric cars to recyclable plastics; there have been several moral responses to the earth's clarion call to caution, many have become a mainstream tech. The development of alternative energy is also bridging the gap between being a moral responsibility and being a much-needed technology that is clean and cost-effective. One of these much-needed technologies is the solar home system.
From electric cars to recyclable plastics; there have been several moral responses to the earth's clarion call to caution, many have become a mainstream tech. The development of alternative energy is also bridging the gap between being a moral responsibility and being a much-needed technology that is clean and cost-effective. One of these much-needed technologies is the solar home system.
 For a long time, I have believed that we need to disrupt education.
For a long time, I have believed that we need to disrupt education.
 Technology is the application of scientific knowledge to the practical aims of human life.
Technology is the application of scientific knowledge to the practical aims of human life.
 The blockchain space is primarily occupied by men, but here are some ways to increase blockchain use among women and level the numbers a bit.
The blockchain space is primarily occupied by men, but here are some ways to increase blockchain use among women and level the numbers a bit.
Software Development](https://hackernoon.com/devops-and-cicd-collaboration-bridging-the-gap-for-efficient-software-development)
 Unlock the power of DevOps and CI/CD collaboration with our easy steps. Streamline software development for efficiency and success.
Unlock the power of DevOps and CI/CD collaboration with our easy steps. Streamline software development for efficiency and success.
 Increasingly sophisticated NFTs are giving rise to a new information dimension with concrete roots in the physical world.
Increasingly sophisticated NFTs are giving rise to a new information dimension with concrete roots in the physical world.
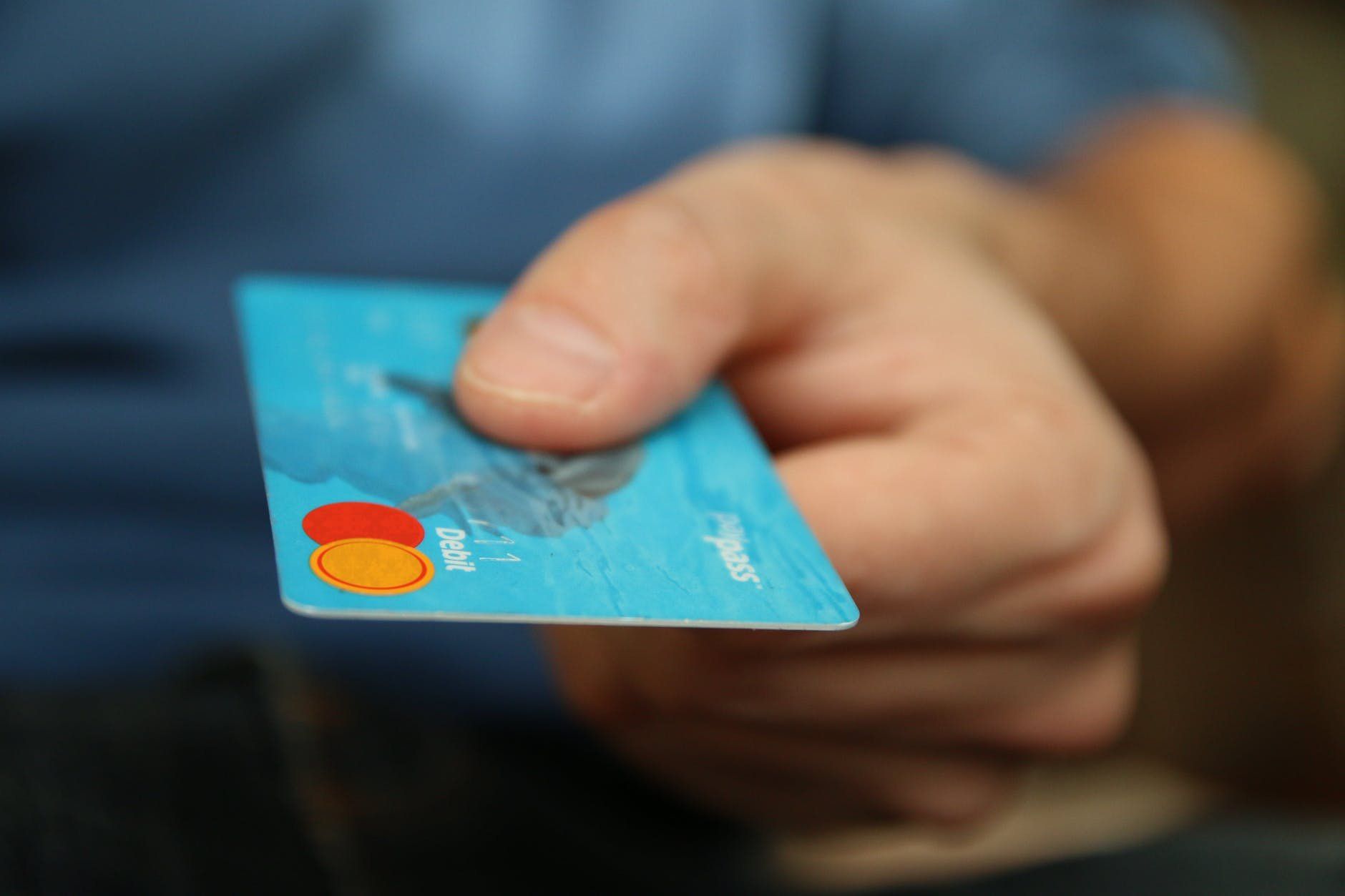 Comparatively to traditional cryptocurrencies, a CBDC is centralized, and thus it is regulated by the issuing organization or country.
Comparatively to traditional cryptocurrencies, a CBDC is centralized, and thus it is regulated by the issuing organization or country.
 I'll show you how to set up a firewall and lock down the whole server for anyone to access with iptables and knockd.
I'll show you how to set up a firewall and lock down the whole server for anyone to access with iptables and knockd.
 A few weeks ago MariaDB launched their new database-as-a-service (DBaaS), SkySQL, amid the Coronavirus Pandemic. While they also offered a $500 credit to get started, as of last week, they announced a program to offer their fully-managed analytics (columnar based storage) service for free to help fight COVID-19.
A few weeks ago MariaDB launched their new database-as-a-service (DBaaS), SkySQL, amid the Coronavirus Pandemic. While they also offered a $500 credit to get started, as of last week, they announced a program to offer their fully-managed analytics (columnar based storage) service for free to help fight COVID-19.
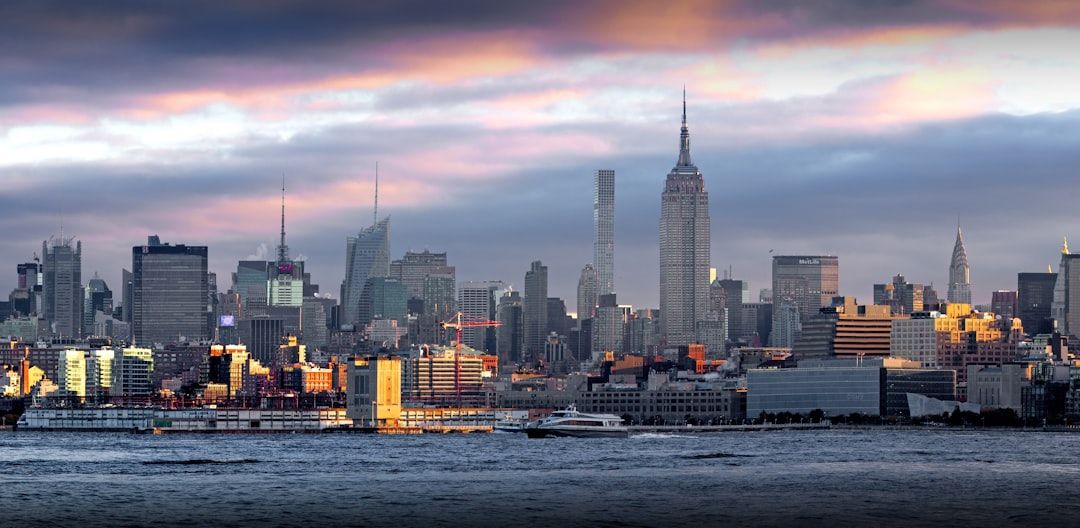 Explore the transformative role of spatial data management in smart cities, addressing its applications, challenges, and benefits.
Explore the transformative role of spatial data management in smart cities, addressing its applications, challenges, and benefits.
 Explore the role of open-source projects w in driving AI innovation. Learn how collaboration and shared knowledge shape AI's future across industries.
Explore the role of open-source projects w in driving AI innovation. Learn how collaboration and shared knowledge shape AI's future across industries.
 Another year, another iPhone with minimal changes. Virtually identical to the 2017 design except for the flat edges, the iPhone 12 that Apple recently announced doesn’t surprise. It pleases, but it doesn’t dazzle. 5G and a series of back magnets, named MagSafe, complete the highlights of a device that will sell well, but that won’t do anything to push the envelope.
Another year, another iPhone with minimal changes. Virtually identical to the 2017 design except for the flat edges, the iPhone 12 that Apple recently announced doesn’t surprise. It pleases, but it doesn’t dazzle. 5G and a series of back magnets, named MagSafe, complete the highlights of a device that will sell well, but that won’t do anything to push the envelope.
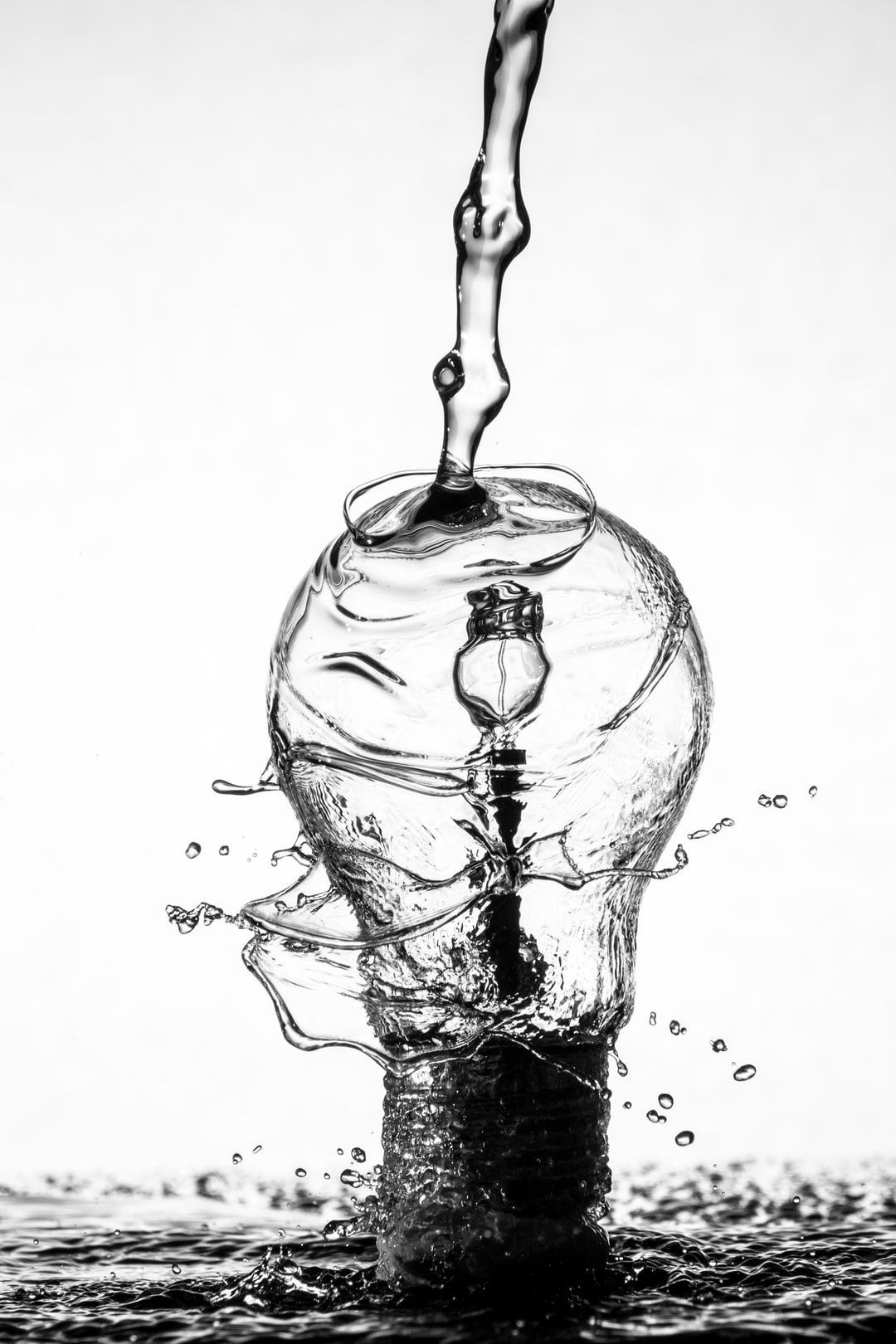 In essence, metaphors are water. The container determines the metaphor’s shape.
In essence, metaphors are water. The container determines the metaphor’s shape.
 Year Highlights for TAIKAI's Product, Community and Challenges.
Year Highlights for TAIKAI's Product, Community and Challenges.
 The new innovations in the robotics industry are creating exciting opportunities for investors.
The new innovations in the robotics industry are creating exciting opportunities for investors.
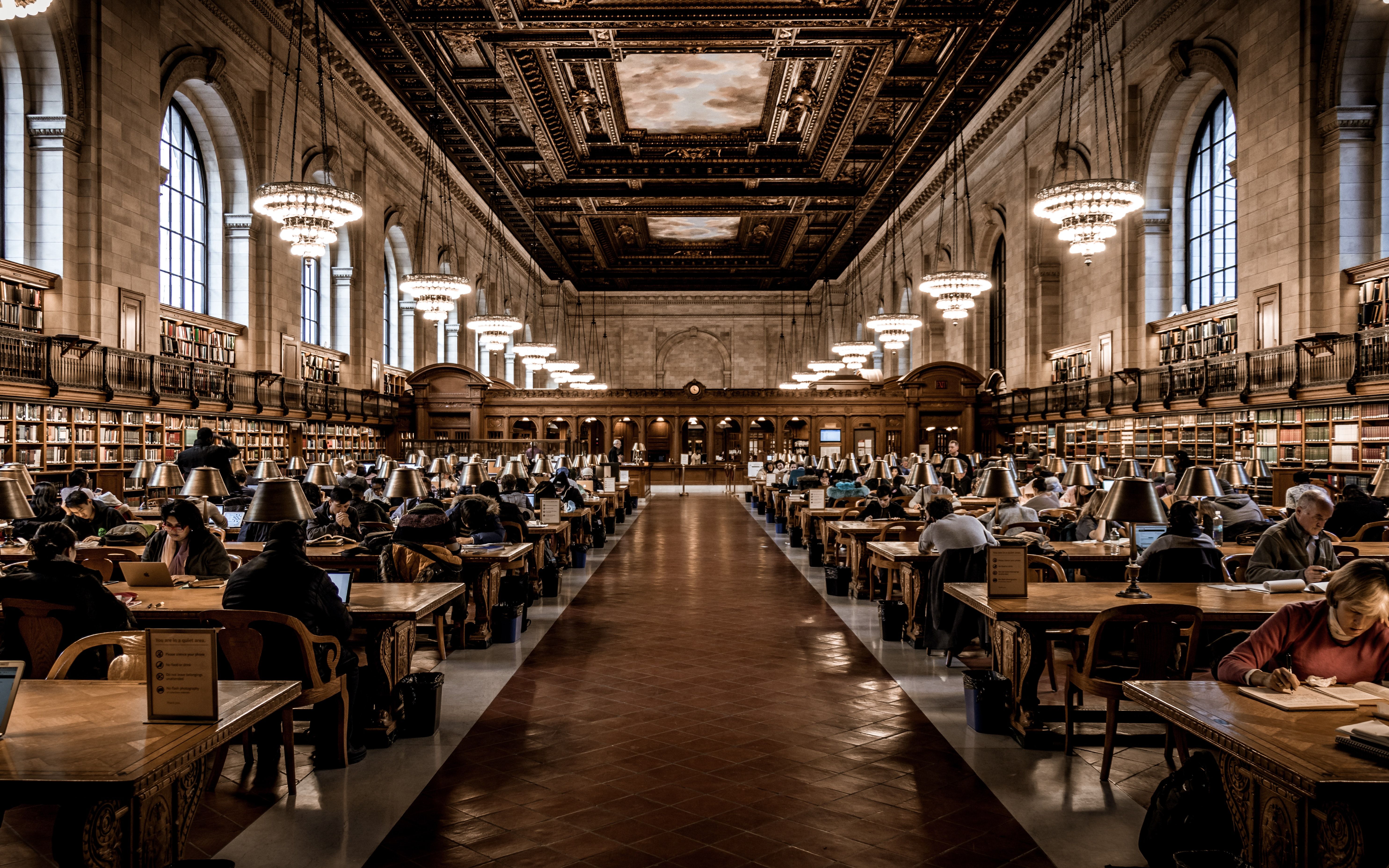 How blockchain implementation in academia would have profound downstream effects in society.
How blockchain implementation in academia would have profound downstream effects in society.
 Exploring the world of brain-computer interface (BCI) technology for disabled creatives and the battle against neurodegenerative disorders.
Exploring the world of brain-computer interface (BCI) technology for disabled creatives and the battle against neurodegenerative disorders.
 Amidst divergent AI governance strategies, global powers navigate cooperation and competition, shaping a future of technological solidarity.
Amidst divergent AI governance strategies, global powers navigate cooperation and competition, shaping a future of technological solidarity.
 EditGAN allows you to control any feature from quick drafts, and it will only edit what you want keeping the rest of the image the same!
EditGAN allows you to control any feature from quick drafts, and it will only edit what you want keeping the rest of the image the same!
 Innovation comes from identifying customer needs and meeting them. It is easy to understand but hard to do. When you start building a new product, you need to m
Innovation comes from identifying customer needs and meeting them. It is easy to understand but hard to do. When you start building a new product, you need to m
 Ever been to meetings you didn't want to be a part of? Meetings you could do better without but, you had to go through thoroughly because you didn't know when your name can be announced and you knew you'd have to respond then. Yeah, we have all been there.
Ever been to meetings you didn't want to be a part of? Meetings you could do better without but, you had to go through thoroughly because you didn't know when your name can be announced and you knew you'd have to respond then. Yeah, we have all been there.
That's where "MSorryWhat!?" - my weekend project comes into play. It's a simple speech-recognition which comes with hot word detection which a user can set, upon complete configuration (which doesn't take more than 5 minutes), MSorryWhat listens to your meeting while you do whatever else it is that you want to do and it notifies you whenever your selected hot word is detected!
It is a simple javascript based application living here. It leverages the Webhooks and Notifications services of IFTTT (If this then that) app, to notify you when your name is announced, on the backend it leverages chrome's speech recognition engine.
I spent 10 minutes to create a demo for the tool which can be viewed here:
 Discover how Michelin-star chef Massimo Bottura's approach can revolutionize your data strategy. Learn to craft unique, impactful data initiatives for your tech
Discover how Michelin-star chef Massimo Bottura's approach can revolutionize your data strategy. Learn to craft unique, impactful data initiatives for your tech
 With Web3 tech and blockchain-based platforms, artists were promised more control over their music and fairer compensation for their work.
With Web3 tech and blockchain-based platforms, artists were promised more control over their music and fairer compensation for their work.
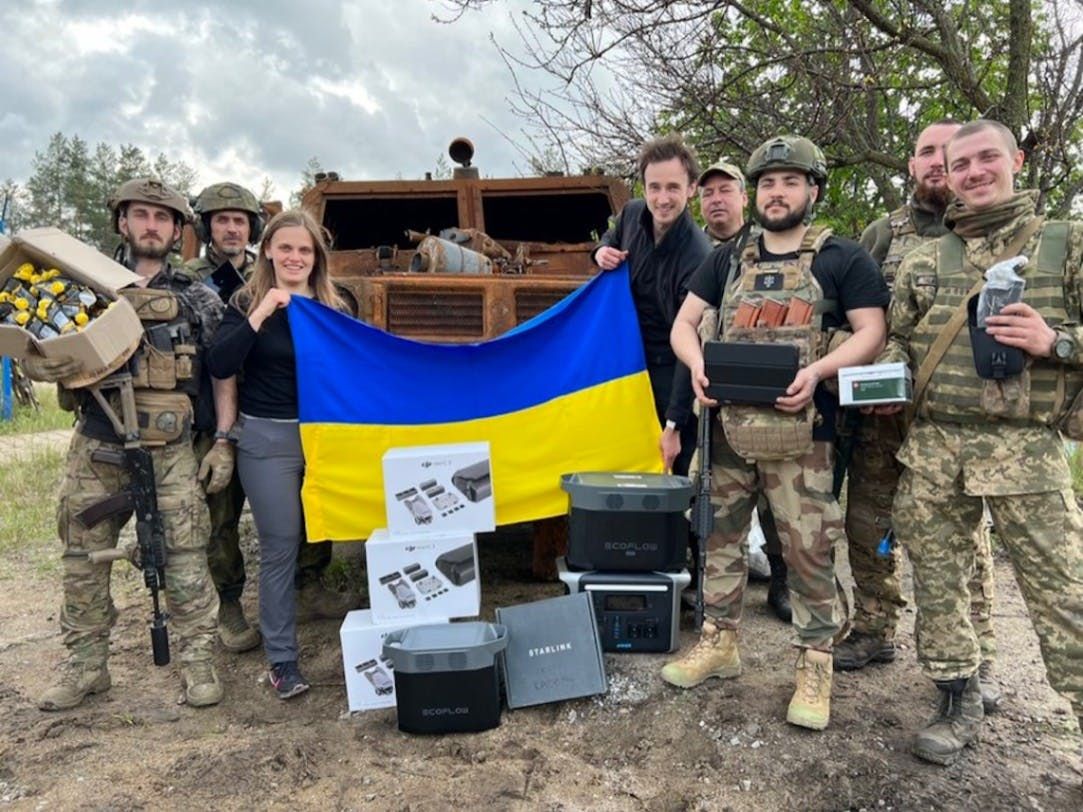 How Dyzga's Paw drove battlefield innovation and volunteer tech supply chains for the Ukrainian army. The Geeks of War.
How Dyzga's Paw drove battlefield innovation and volunteer tech supply chains for the Ukrainian army. The Geeks of War.
 Learn an 8-step framework for building sustainable tech startups that balance profitability, customer value, and ethical workforce practices for lasting growth.
Learn an 8-step framework for building sustainable tech startups that balance profitability, customer value, and ethical workforce practices for lasting growth.
 Daniel Jeffries is Chief Technology Evangelist at Pachyderm, author, engineer, futurist and pro blogger.
Daniel Jeffries is Chief Technology Evangelist at Pachyderm, author, engineer, futurist and pro blogger.
 Startups: It’s not about being first, it’s about playing well with others. Open Innovation = sharing ideas, collaborating, and winning together.
Startups: It’s not about being first, it’s about playing well with others. Open Innovation = sharing ideas, collaborating, and winning together.
 AI is taking on an increasingly important role in modern society, so it is critical that algorithms can understand and utilize many different languages.
AI is taking on an increasingly important role in modern society, so it is critical that algorithms can understand and utilize many different languages.
 For some time now I’ve been making an unassailable and unchallenged argument that Bitcoin and Bitcoin services are no different to the monies and services found in video games. Now, thanks to my favourite gamer, we have one of the best examples yet of “Bitcoin Doublethink”, second only to the very cute Pigsby which uses a “Blockchain” for its database.
For some time now I’ve been making an unassailable and unchallenged argument that Bitcoin and Bitcoin services are no different to the monies and services found in video games. Now, thanks to my favourite gamer, we have one of the best examples yet of “Bitcoin Doublethink”, second only to the very cute Pigsby which uses a “Blockchain” for its database.
 WiFi Map shows you more than just public hotspots that are available nearby, but provides great time-saving help by also offering passwords for these hotspots.
WiFi Map shows you more than just public hotspots that are available nearby, but provides great time-saving help by also offering passwords for these hotspots.
 Told to chase grades and play it safe, I chose to build AI and quantum tools instead—without a roadmap, rewards, or permission. Here’s why I never stopped.
Told to chase grades and play it safe, I chose to build AI and quantum tools instead—without a roadmap, rewards, or permission. Here’s why I never stopped.
 Creativity goes hand-in-hand with innovation, and it’s hardly possible to imagine a startup without a spark of creative thought. But what if there are processes that are stifling your team’s full creative potential, making it harder to work, develop, and stay ahead of the competition? What are the key elements that inspire creativity within a team, and how can team leaders help? This practical guide summarizes the most effective methods, as well as mistakes to watch out for.
Creativity goes hand-in-hand with innovation, and it’s hardly possible to imagine a startup without a spark of creative thought. But what if there are processes that are stifling your team’s full creative potential, making it harder to work, develop, and stay ahead of the competition? What are the key elements that inspire creativity within a team, and how can team leaders help? This practical guide summarizes the most effective methods, as well as mistakes to watch out for.
 Learn about the pursuit to solve one of the most severe and long-prevailing problems in fintech.
Learn about the pursuit to solve one of the most severe and long-prevailing problems in fintech.
 The Red Queen Effect: Thriving in a fast-paced world. Insights on adaptation, innovation, and growth in business & life.
The Red Queen Effect: Thriving in a fast-paced world. Insights on adaptation, innovation, and growth in business & life.
 No, seriously, why not? It’s cool to be a designer, a creative person who DJs vinyl sets in a funky club at night and starts the morning with a double-filter coffee and a vegan croissant overlooking the sea to get inspired for the next advertising campaign. Not a bad stereotype, right?
No, seriously, why not? It’s cool to be a designer, a creative person who DJs vinyl sets in a funky club at night and starts the morning with a double-filter coffee and a vegan croissant overlooking the sea to get inspired for the next advertising campaign. Not a bad stereotype, right?
 FIFA teams up with Modex for FIFA+ Collect, offering exclusive digital collectibles & 2026 World Cup Final™ tickets.
FIFA teams up with Modex for FIFA+ Collect, offering exclusive digital collectibles & 2026 World Cup Final™ tickets.
 Can we learn through video games? Can eLearning games completely replace textbooks? What makes us have fun?
Can we learn through video games? Can eLearning games completely replace textbooks? What makes us have fun?
 My generation has seen a world much less forgiving. We grew up understanding the fundamentals of climate change, and saw rising sea levels and record setting temperatures all over the world.
My generation has seen a world much less forgiving. We grew up understanding the fundamentals of climate change, and saw rising sea levels and record setting temperatures all over the world.
 TAIKAI, a Startup of the Year nominee, helps organizations run their hackathons and innovation challenges, acting as an online management tool.
TAIKAI, a Startup of the Year nominee, helps organizations run their hackathons and innovation challenges, acting as an online management tool.
 Ignatica CEO and Co-Founder Manuel San Miguel discusses why he founded the company, AI and Machine Learning and why he should have gone to Japan sooner.
Ignatica CEO and Co-Founder Manuel San Miguel discusses why he founded the company, AI and Machine Learning and why he should have gone to Japan sooner.
 About twenty times in my career as a Computer Scientist and Public Inventor I have somehow informally presented an idea to someone or a group of people, only to hear them say, “That’s been done.”
About twenty times in my career as a Computer Scientist and Public Inventor I have somehow informally presented an idea to someone or a group of people, only to hear them say, “That’s been done.”
 How Technologies of Ambition Shape Society's Evolution
How Technologies of Ambition Shape Society's Evolution
 Disruption is the name of the game these days, from current remote work settings due to COVID-19 to even more normal, every-day disruptions (like the process of moving and renovating a home, which I’m currently experiencing first-hand). As a habit-driven developer myself, I have come to the realization that the stable work environment I’ve come to rely on is no more.
Disruption is the name of the game these days, from current remote work settings due to COVID-19 to even more normal, every-day disruptions (like the process of moving and renovating a home, which I’m currently experiencing first-hand). As a habit-driven developer myself, I have come to the realization that the stable work environment I’ve come to rely on is no more.
 In the classic “diffusion of innovations” theory, new technology has to progress from innovators to early adopters before it can go mainstream.
In the classic “diffusion of innovations” theory, new technology has to progress from innovators to early adopters before it can go mainstream.
 Stay ahead in AI with these 8 essential newsletters for founders. Get actionable insights, practical advice, and real-world applications to build smarter.
Stay ahead in AI with these 8 essential newsletters for founders. Get actionable insights, practical advice, and real-world applications to build smarter.
 What makes some ideas change the world? The simple answer is it's about turning your ideas into real solutions. Not enough, isn’t it? Okay then, keep reading.
What makes some ideas change the world? The simple answer is it's about turning your ideas into real solutions. Not enough, isn’t it? Okay then, keep reading.
 These 7 Startup Founders used the challenges presented by the Covid-19 pandemic to propel them forward. From asynchronous communication apps to a self-sanitizin
These 7 Startup Founders used the challenges presented by the Covid-19 pandemic to propel them forward. From asynchronous communication apps to a self-sanitizin
 In 2019, the value of U.S. venture capital investments was $108 billion. In the same year, the federal government spent $131 billion on research and development.
In 2019, the value of U.S. venture capital investments was $108 billion. In the same year, the federal government spent $131 billion on research and development.
 Hola! Örvendek! Gutentag! Kayfak? In my neverending search to connect with other humans, I’ve always been fond of learning new languages…
Hola! Örvendek! Gutentag! Kayfak? In my neverending search to connect with other humans, I’ve always been fond of learning new languages…
 Innovating for inclusivity in tech startups: How solving for edge cases leads to better solutions for all users, illustrated through legal tech advancements.
Innovating for inclusivity in tech startups: How solving for edge cases leads to better solutions for all users, illustrated through legal tech advancements.
 The same tech that can built dystopian futures can help support humans in real growth, education, equality, sense-making and wise decision making.
The same tech that can built dystopian futures can help support humans in real growth, education, equality, sense-making and wise decision making.
 Overly-strict or rigid regulatory frameworks can carry significant risks, including stifling innovation and progress.
Overly-strict or rigid regulatory frameworks can carry significant risks, including stifling innovation and progress.
 In this story, we will explain the true potential of blockchain and why it's a lot more interesting to use this technology outside of cryptocurrency.
In this story, we will explain the true potential of blockchain and why it's a lot more interesting to use this technology outside of cryptocurrency.
 AI teams ship fast, but users struggle to keep up. Learn why rapid iteration creates trust loss and how to close the Velocity–Comprehension Gap.
AI teams ship fast, but users struggle to keep up. Learn why rapid iteration creates trust loss and how to close the Velocity–Comprehension Gap.
 LLM-powered daily newsletters stuck repeating content? Learn why RAG stops too early and how local caching creates unique, diverse outputs every day.
LLM-powered daily newsletters stuck repeating content? Learn why RAG stops too early and how local caching creates unique, diverse outputs every day.
 Looking to ride a business venture into the unknown? Learn the 5 steps of frontier pioneering first!
Looking to ride a business venture into the unknown? Learn the 5 steps of frontier pioneering first!
 When considering where to launch their startup, many entrepreneurs naturally think of the “Innovation Hubs” of Boston, the San Francisco Bay Area, and New York City. After all, that is where the deals are, right? Well, they’re not the only places where companies are being launched and receiving game-changing investments.
When considering where to launch their startup, many entrepreneurs naturally think of the “Innovation Hubs” of Boston, the San Francisco Bay Area, and New York City. After all, that is where the deals are, right? Well, they’re not the only places where companies are being launched and receiving game-changing investments.
 Without the CDN infrastructure, Disney+'s 100 million subscribers will not be able to enjoy the movies at home during the pandemic.
Without the CDN infrastructure, Disney+'s 100 million subscribers will not be able to enjoy the movies at home during the pandemic.
 What is the future of retail in the post-COVID-19 world? The 2021-reality trends show how technology and innovations help retailers face challenges.
What is the future of retail in the post-COVID-19 world? The 2021-reality trends show how technology and innovations help retailers face challenges.
 Monopoly is a word you hear probably once a month or so, maybe once a week if you’re a big board games fan. The word itself has stopped being associated with terrible business practices and is now considered more as honest fun with your friends.
Monopoly is a word you hear probably once a month or so, maybe once a week if you’re a big board games fan. The word itself has stopped being associated with terrible business practices and is now considered more as honest fun with your friends.
 Explore the essentials of fostering innovation in startups—cultivating creativity, embracing risk, and building a culture of ingenuity.
Explore the essentials of fostering innovation in startups—cultivating creativity, embracing risk, and building a culture of ingenuity.
 America's critical infrastructure runs on GPS — but as threats grow, officials warn it's time to build resilient alternatives before disaster strikes.
America's critical infrastructure runs on GPS — but as threats grow, officials warn it's time to build resilient alternatives before disaster strikes.
 We named our startup InvoZone (A zone full of innovative people). leading innovators in web and mobile development using AI, IoT, blockchain.
We named our startup InvoZone (A zone full of innovative people). leading innovators in web and mobile development using AI, IoT, blockchain.
 Explore the intricate balance between innovation and maintenance in software development. Uncover strategies to ensure growth while maintaining product integrit
Explore the intricate balance between innovation and maintenance in software development. Uncover strategies to ensure growth while maintaining product integrit
 Varun Shah is a thought leader in the IT space. He has worked for Sony PlayStation, Medimpact Healthcare Systems, and Amazon.
Varun Shah is a thought leader in the IT space. He has worked for Sony PlayStation, Medimpact Healthcare Systems, and Amazon.

 Feature Image: HackerNoon’s Stable Diffusion AI, Prompt” Dealer’s choice”
Feature Image: HackerNoon’s Stable Diffusion AI, Prompt” Dealer’s choice”
 Organizations of all sizes treat data security and regulatory compliance like a roadblock instead of a design consideration. Over the last ten years, I have worked with scores of organizations to modernize and improve their software and data practices. Information security is a challenging position, as it is the job of security professionals to prevent companies from making changes that could seriously damage reputation and revenue or breach compliance and incur significant penalties. There are two general archetypes of security teams that I have interfaced with:
Organizations of all sizes treat data security and regulatory compliance like a roadblock instead of a design consideration. Over the last ten years, I have worked with scores of organizations to modernize and improve their software and data practices. Information security is a challenging position, as it is the job of security professionals to prevent companies from making changes that could seriously damage reputation and revenue or breach compliance and incur significant penalties. There are two general archetypes of security teams that I have interfaced with:
 Gen Z is a generation of young people who are interested in improving their lives. They can benefit from informal education if they do not choose college
Gen Z is a generation of young people who are interested in improving their lives. They can benefit from informal education if they do not choose college
 Computer vision applications have become ubiquitous nowadays. It’s hard to think of a domain where the ability of computers to “see” what’s going on around them has not yet been leveraged.
Computer vision applications have become ubiquitous nowadays. It’s hard to think of a domain where the ability of computers to “see” what’s going on around them has not yet been leveraged.
 From network effects to infrastructure costs, discover why the decentralized internet faces insurmountable challenges despite years of promises
From network effects to infrastructure costs, discover why the decentralized internet faces insurmountable challenges despite years of promises
 On what it means to be an orchestrator and why the more i get into tech, the more i want to go analog.
On what it means to be an orchestrator and why the more i get into tech, the more i want to go analog.
 Perhaps dating back to the beginning of humankind, we have reached out to touch someone. Not just one of the best advertising slogans of modern history, handshakes, hugs, and kisses are ways we touch someone to convey welcome, partnership, love, friendship, empathy, acceptance, and countless other emotions with acquaintances and loved ones. With the threat of virus transmissions now at the forefront of people’s minds, is there a place for the human touch in our future?
Perhaps dating back to the beginning of humankind, we have reached out to touch someone. Not just one of the best advertising slogans of modern history, handshakes, hugs, and kisses are ways we touch someone to convey welcome, partnership, love, friendship, empathy, acceptance, and countless other emotions with acquaintances and loved ones. With the threat of virus transmissions now at the forefront of people’s minds, is there a place for the human touch in our future?
 The purpose of this article isn't investment advice. It isn't to sell a product. However, it is to sell a vision. I am writing about my startup known as Stark Drones that is working on some quite innovative technologies. In the age of copycat products and the whole Silicon Valley bugs, comes some startups wanting to build truly disruptive technologies.
The purpose of this article isn't investment advice. It isn't to sell a product. However, it is to sell a vision. I am writing about my startup known as Stark Drones that is working on some quite innovative technologies. In the age of copycat products and the whole Silicon Valley bugs, comes some startups wanting to build truly disruptive technologies.
 Crises lead to behavioral change, both short-term and longer term. Today’s temporary response to a disruption can become tomorrow’s new norms or even habits. Those behavioral changes can cast a spotlight on technological solutions that help people cope with change. And we already know of many occurring during the COVID-19 pandemic.
Crises lead to behavioral change, both short-term and longer term. Today’s temporary response to a disruption can become tomorrow’s new norms or even habits. Those behavioral changes can cast a spotlight on technological solutions that help people cope with change. And we already know of many occurring during the COVID-19 pandemic.
 Lama, the regulated cryptocurrency exchange, has unveiled Visa cards for its users.
Lama, the regulated cryptocurrency exchange, has unveiled Visa cards for its users.
 Both open-source software and hackathons encourage collaboration between a number of skilled individuals, often rallying them around a single goal.
Both open-source software and hackathons encourage collaboration between a number of skilled individuals, often rallying them around a single goal.
 UCROWDME is the world’s first idea currency and global ideas marketplace where people can sell their ideas to businesses, protected on the blockchain.
UCROWDME is the world’s first idea currency and global ideas marketplace where people can sell their ideas to businesses, protected on the blockchain.
 As you prepare for your board meetings, you might be struggling to find some good news to share with the board. Sure, it’s a mistake to go into a board meeting pretending everything is going well — board meetings are a valuable opportunity to share your startup’s challenges and get guidance from a panel of trusted advisors.
As you prepare for your board meetings, you might be struggling to find some good news to share with the board. Sure, it’s a mistake to go into a board meeting pretending everything is going well — board meetings are a valuable opportunity to share your startup’s challenges and get guidance from a panel of trusted advisors.
 Unlock the true potential of enterprise digital transformation with insights into common pitfalls and signs for an upgrade.
Unlock the true potential of enterprise digital transformation with insights into common pitfalls and signs for an upgrade.
 Futureproofing is at the forefront of every entrepreneur’s mind. But putting ideas into practice can be a different challenge altogether.
Futureproofing is at the forefront of every entrepreneur’s mind. But putting ideas into practice can be a different challenge altogether.
 London has always been a world leader in innovation. From the invention of the first mobile phone, to the creation of a ‘metro’ system that's now used worldwide, London has continually risen as an international tech hub.
London has always been a world leader in innovation. From the invention of the first mobile phone, to the creation of a ‘metro’ system that's now used worldwide, London has continually risen as an international tech hub.
 Attributing for one-fifth of the planet's population, China is undeniably an economic superpower that's empowered to become a global blockchain leader in terms of adoption, utility, and overall innovation.
Attributing for one-fifth of the planet's population, China is undeniably an economic superpower that's empowered to become a global blockchain leader in terms of adoption, utility, and overall innovation.
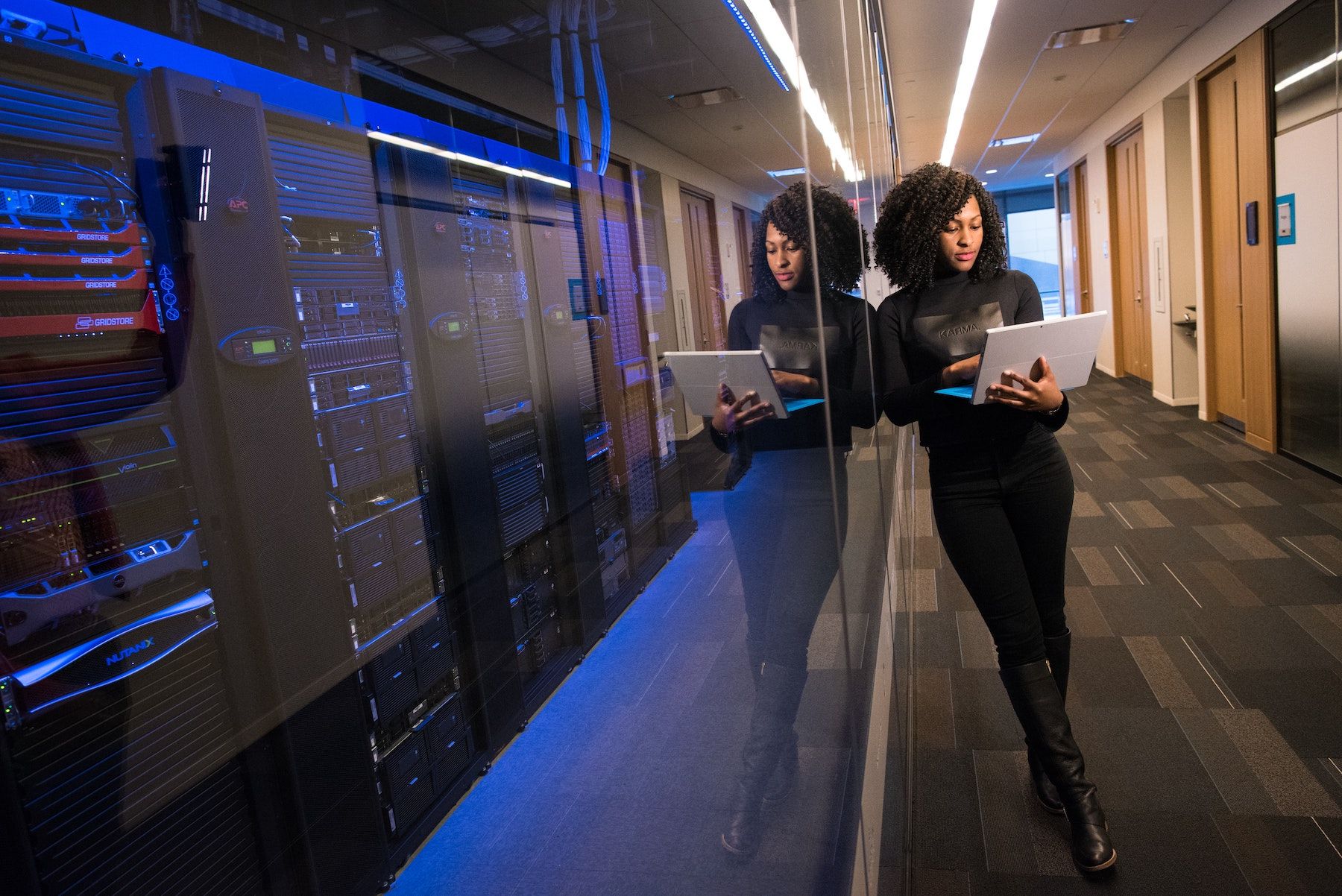 In the healthcare landscape, providers and lawmakers alike are faced with the challenge of making the best possible decisions for patients and the industry as a whole. From choosing the best treatments to using resources in a responsible manner, medical leaders are making decisions on a daily basis that can significantly impact health outcomes and costs.
In the healthcare landscape, providers and lawmakers alike are faced with the challenge of making the best possible decisions for patients and the industry as a whole. From choosing the best treatments to using resources in a responsible manner, medical leaders are making decisions on a daily basis that can significantly impact health outcomes and costs.
 Crypto startups used to raise millions on hype, but most failed fast. Learn why real product-market fit, not token sales, defines success in 2025.
Crypto startups used to raise millions on hype, but most failed fast. Learn why real product-market fit, not token sales, defines success in 2025.
 Unlike GenAI and AR dresses, some wearable and outdoor tech innovations may not appear on the catwalks but could be essential (and save your life)
Unlike GenAI and AR dresses, some wearable and outdoor tech innovations may not appear on the catwalks but could be essential (and save your life)
 U.S. passes the GENIUS Act, creating clear stablecoin rules and boosting innovation, while Canada risks losing fintech talent amid regulatory delays.
U.S. passes the GENIUS Act, creating clear stablecoin rules and boosting innovation, while Canada risks losing fintech talent amid regulatory delays.
 The founders of AnavClouds Software are both Salesforce professionals, doing well in their own spheres of work. They happened to meet through a common friend of theirs and started working together for some time. As the volumes grew they decided over a cup of steaming coffee that it was the right time to start a venture of their own, in 2016.
The founders of AnavClouds Software are both Salesforce professionals, doing well in their own spheres of work. They happened to meet through a common friend of theirs and started working together for some time. As the volumes grew they decided over a cup of steaming coffee that it was the right time to start a venture of their own, in 2016.
 Facial recognition is an upcoming feature. We see it in marketing and in our devices. In this slogging thread, we discussed the pros and cons of this feature.
Facial recognition is an upcoming feature. We see it in marketing and in our devices. In this slogging thread, we discussed the pros and cons of this feature.
 A slow load time on your devices down the borderline unacceptable in today’s society. The average consumer now expects everything to work instantly.
A slow load time on your devices down the borderline unacceptable in today’s society. The average consumer now expects everything to work instantly.
 Would Wonder Woman 1984 have worked better as Wonder Woman 1994 or Wonder Woman 2004?
Would Wonder Woman 1984 have worked better as Wonder Woman 1994 or Wonder Woman 2004?
 We are currently standing at the helm of the fourth revolution, where advances in computing power and lower costs of computing has enabled us to apply natural language processing, artificial intelligence, machine learning and neural networks to seemingly mundane tasks. Though our previous generations were instrumental in developing efficiencies in established processes, technologies of today go much beyond – offering an opportunity to augment these in new ways, and not simply eradicate established methods of managing wealth. These opportunities for wealth management are genuinely path breaking.
We are currently standing at the helm of the fourth revolution, where advances in computing power and lower costs of computing has enabled us to apply natural language processing, artificial intelligence, machine learning and neural networks to seemingly mundane tasks. Though our previous generations were instrumental in developing efficiencies in established processes, technologies of today go much beyond – offering an opportunity to augment these in new ways, and not simply eradicate established methods of managing wealth. These opportunities for wealth management are genuinely path breaking.
 Take your worries away thesis couture is here to save the day. A Stiletto made by several professionals and engineers to lessen the pain of women's footwear.
Take your worries away thesis couture is here to save the day. A Stiletto made by several professionals and engineers to lessen the pain of women's footwear.
 There are a lot of unhappy developers and this shouldn't be the case.
There are a lot of unhappy developers and this shouldn't be the case.
](https://hackernoon.com/vibe-coding-lowering-the-cost-of-trying)
 The hidden value of vibe coding isn’t just speed; it’s about increasing the pace of progress.
The hidden value of vibe coding isn’t just speed; it’s about increasing the pace of progress.
 The landscape of complex IT projects demands not only technical expertise but also a mastery of managing innovations. How to adjust to these challenges?
The landscape of complex IT projects demands not only technical expertise but also a mastery of managing innovations. How to adjust to these challenges?
 Automation is an exciting prospect. Who doesn’t like the idea of having menial tasks completed quicker and more effectively than they could have been by a human?
Automation is an exciting prospect. Who doesn’t like the idea of having menial tasks completed quicker and more effectively than they could have been by a human?
 The future of financial services won't be built in silos—it will be shaped in ecosystems.
The future of financial services won't be built in silos—it will be shaped in ecosystems.
 Discover how tech organizations thrive by prioritizing principles, trust, innovation, and learning in the dynamic digital landscape. Adapt or fade.
Discover how tech organizations thrive by prioritizing principles, trust, innovation, and learning in the dynamic digital landscape. Adapt or fade.
 Tech Companies are running internal hackathons to design new features, build new products, and increase employee and talent engagement.
Tech Companies are running internal hackathons to design new features, build new products, and increase employee and talent engagement.
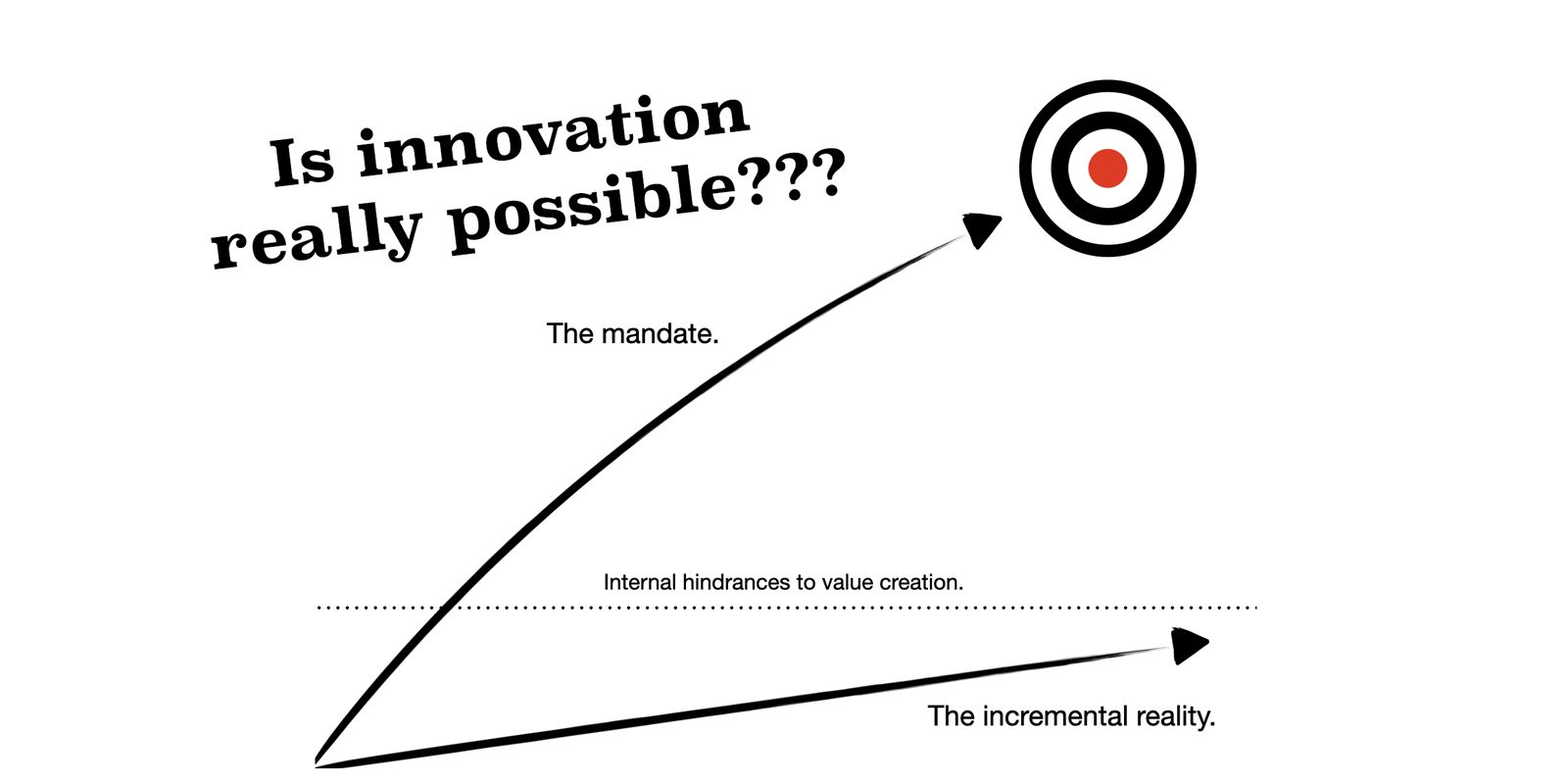 While serious resources can be spent on creative vision, rarely is anyone given the tools to properly test an idea in-market quickly at high fidelity.
While serious resources can be spent on creative vision, rarely is anyone given the tools to properly test an idea in-market quickly at high fidelity.
 Learn more about Porto-based startup nBanks in this interview with nBanks CEO Orlando Costa.
Learn more about Porto-based startup nBanks in this interview with nBanks CEO Orlando Costa.
 Syed Aamir Aarfi integrates AI/ML to enhance e-commerce, SaaS, and supply chains, driving innovation with a customer-centered approach.
Syed Aamir Aarfi integrates AI/ML to enhance e-commerce, SaaS, and supply chains, driving innovation with a customer-centered approach.
 Serverless doesn’t mean operating an application without a server, it means that you can operate the app without having to manage a server.
Serverless doesn’t mean operating an application without a server, it means that you can operate the app without having to manage a server.
 Many people wonder what the point of open source is? For those who don't get it, they think it is giving stuff out for free. They don't understand how BeagleBone exploded the way they did. More importantly, they don't seem to understand that the internet as we know it, or many innovations in terms of the modern world, wouldn't exist or be the same if it wasn't for open source. Even when you look at proprietary tech companies like Apple, one can argue figures such as Steve Wozniak were greatly inspired by the open hardware movement many decades ago.
Many people wonder what the point of open source is? For those who don't get it, they think it is giving stuff out for free. They don't understand how BeagleBone exploded the way they did. More importantly, they don't seem to understand that the internet as we know it, or many innovations in terms of the modern world, wouldn't exist or be the same if it wasn't for open source. Even when you look at proprietary tech companies like Apple, one can argue figures such as Steve Wozniak were greatly inspired by the open hardware movement many decades ago.
 Remember when we mere mortals were able to cognitively perceive and understand the pace at which innovations were made?
Remember when we mere mortals were able to cognitively perceive and understand the pace at which innovations were made?
Me neither - that kind of linear development model sounds like something that might've held true during the Industrial Revolution.
 Many of us have heard the phrase "content is king" when discussing online marketing. Although the expression was originally coined back in 1996 by Microsoft founder, Bill Gates, it has never been more relevant than it is today. The phrase is rooted in the idea that valuable, engaging, and exciting material is essential for success in the Ecommerce space.
Many of us have heard the phrase "content is king" when discussing online marketing. Although the expression was originally coined back in 1996 by Microsoft founder, Bill Gates, it has never been more relevant than it is today. The phrase is rooted in the idea that valuable, engaging, and exciting material is essential for success in the Ecommerce space.
 From biotechnology to artificial intelligence (AI), innovation is keeping pace with our ability to comprehend its effects.
From biotechnology to artificial intelligence (AI), innovation is keeping pace with our ability to comprehend its effects.
 Discover how Solution Architect Ashish Komal leverages 16+ years of expertise in cloud migration to drive innovation, tackle challenges, and inspire change.
Discover how Solution Architect Ashish Komal leverages 16+ years of expertise in cloud migration to drive innovation, tackle challenges, and inspire change.
with Open Innovation Challenges](https://hackernoon.com/how-to-capture-disruptive-ideas-with-open-innovation-challenges-sn3c3355)
 Henry Chesbrough, a professor at the University of California Berkeley Haas School of Business, coined the term “open innovation” almost 2 decades ago.
Henry Chesbrough, a professor at the University of California Berkeley Haas School of Business, coined the term “open innovation” almost 2 decades ago.
 Join us in our chat with Jason Labaw, the CEO and founder of Social Bee, an augmented reality platform that brings the tourism experience to the next level.
Join us in our chat with Jason Labaw, the CEO and founder of Social Bee, an augmented reality platform that brings the tourism experience to the next level.
 Every startup in its early stages is going to think about building brand awareness, why? Well, brand awareness is an essential tool that is going to let the world know about your brand and the work you do. Sometimes this is all that you need when you are working on your startup because having a proper brand is going to mean that you get recognized easily and you get more revenue.
Every startup in its early stages is going to think about building brand awareness, why? Well, brand awareness is an essential tool that is going to let the world know about your brand and the work you do. Sometimes this is all that you need when you are working on your startup because having a proper brand is going to mean that you get recognized easily and you get more revenue.
 Insights on digital banking transformation, strategies, and challenges from expert Alex Babin, focusing on innovation and agile methods in fintech.
Insights on digital banking transformation, strategies, and challenges from expert Alex Babin, focusing on innovation and agile methods in fintech.
 It’s absolutely mind-boggling how much time and money we invest in combating cybercrime when we could be channeling those resources towards new innovations
It’s absolutely mind-boggling how much time and money we invest in combating cybercrime when we could be channeling those resources towards new innovations
 My appeal though simple is significant: can you please, pretty please, create a noise-cancelling device specifically designed for snoring partners?
My appeal though simple is significant: can you please, pretty please, create a noise-cancelling device specifically designed for snoring partners?
 Learn how to invest into crypto safely. The world's first diversified crypto fund. Investor tips. Business tips.
Learn how to invest into crypto safely. The world's first diversified crypto fund. Investor tips. Business tips.
 Digital transformation is not one single thing to implement, it is a core alignment with continuous investment in innovation and excellence.
Digital transformation is not one single thing to implement, it is a core alignment with continuous investment in innovation and excellence.
 It’s really exciting how the technology we use to do our work and make our lives easier just keeps getting better. I’m pleased that Siri and Alexa know me on a first-name basis. Amazon can often predict what program I want to watch, a book I’d like to read, or clothes I’d like to buy. Roomba has figured out just the right places to vacuum, and I simply have to pick up my iPhone and it can recognize me automatically. Now that’s great service!
It’s really exciting how the technology we use to do our work and make our lives easier just keeps getting better. I’m pleased that Siri and Alexa know me on a first-name basis. Amazon can often predict what program I want to watch, a book I’d like to read, or clothes I’d like to buy. Roomba has figured out just the right places to vacuum, and I simply have to pick up my iPhone and it can recognize me automatically. Now that’s great service!
 Crowdfunding and online charity platforms have exploded, allowing individuals, charities, and entrepreneurs to fundraise from a large pool of potential donors.
Crowdfunding and online charity platforms have exploded, allowing individuals, charities, and entrepreneurs to fundraise from a large pool of potential donors.
 Last Thursday, I was lucky to interview Warren Schirtzinger during a live webinar, one of the co-creators of the “Chasm” framework in the 80s, later polished and popularized by Geoffrey Moore in the book “Crossing the Chasm”.
Last Thursday, I was lucky to interview Warren Schirtzinger during a live webinar, one of the co-creators of the “Chasm” framework in the 80s, later polished and popularized by Geoffrey Moore in the book “Crossing the Chasm”.
 Satellite imagery is not new. It has been around since 1960. What has changed is the way we process those images.
Satellite imagery is not new. It has been around since 1960. What has changed is the way we process those images.
 Our main success metric is the satisfaction of our team. If our team is doing well and happy, the customers will be too and the company will continue to grow.
Our main success metric is the satisfaction of our team. If our team is doing well and happy, the customers will be too and the company will continue to grow.
 Artificial Intelligence (AI) is radically transforming businesses across the world.
Businesses and people must adapt quickly or risk becoming irrelevant.
Artificial Intelligence (AI) is radically transforming businesses across the world.
Businesses and people must adapt quickly or risk becoming irrelevant.
 Augmented Reality technology can improve customer experience, thus creating an environment of interaction between agents and clients.
Augmented Reality technology can improve customer experience, thus creating an environment of interaction between agents and clients.
 Solving big problems requires new paradigms, not old tools—history shows breakthroughs come from shifting perspectives, not more information.
Solving big problems requires new paradigms, not old tools—history shows breakthroughs come from shifting perspectives, not more information.
 The companies that choose to invest in AR smart glasses make it possible to have a long-term position in the market by leading the field in which they operate.
The companies that choose to invest in AR smart glasses make it possible to have a long-term position in the market by leading the field in which they operate.
 Exploring African Entrepreneurship: Examining the why, education and mindset shift for successful businesses amidst a growing population
Exploring African Entrepreneurship: Examining the why, education and mindset shift for successful businesses amidst a growing population
 Do research, identify a market gap and create a product to fill it: the classic entrepreneur’s journey.
Do research, identify a market gap and create a product to fill it: the classic entrepreneur’s journey.
 In the 20th century, scientists and thinkers predicted that technology would change our lives fundamentally. Despite our technological achievements, these changes have not materialized.
In the 20th century, scientists and thinkers predicted that technology would change our lives fundamentally. Despite our technological achievements, these changes have not materialized.
 Step One: Prospecting
Step One: Prospecting
 7/10/2023: Top 5 stories on the Hackernoon homepage!
7/10/2023: Top 5 stories on the Hackernoon homepage!
 This article series tries to put forward General Concepts, Principles and Guidelines for Innovative Software Product Design and Development
This article series tries to put forward General Concepts, Principles and Guidelines for Innovative Software Product Design and Development
 In his essay, Fred Wilson wrote in AVC blog about technical innovation, business model innovation, and the last two big waves we had in the tech industry:
In his essay, Fred Wilson wrote in AVC blog about technical innovation, business model innovation, and the last two big waves we had in the tech industry:
 Should governments regulate open foundation models? If yes, how? That's the key to finding a balance between innovation and regulation. Case in point: EU AI Act
Should governments regulate open foundation models? If yes, how? That's the key to finding a balance between innovation and regulation. Case in point: EU AI Act
 “In these unprecedented times…” People write unprecedentedly good Hacker Noon posts. One such writer is David Deal from the United States - double Noonie Nominee in the Back the Internet and Future Heroes Award Categories. Scroll down for David's take on tech today.
“In these unprecedented times…” People write unprecedentedly good Hacker Noon posts. One such writer is David Deal from the United States - double Noonie Nominee in the Back the Internet and Future Heroes Award Categories. Scroll down for David's take on tech today.
 I want to put forward the idea of software recycle. Software recycle is software reuse taken to a new level.
I want to put forward the idea of software recycle. Software recycle is software reuse taken to a new level.
 Under Elon Musk's leadership, X has become a bold, experimental and innovative platform by mastering 5 fundamentals of product management.
Under Elon Musk's leadership, X has become a bold, experimental and innovative platform by mastering 5 fundamentals of product management.
 Elon with his genius intellect can help us have an internet where we can freely join forums and online groups without the fear of dealing with the crypto shills
Elon with his genius intellect can help us have an internet where we can freely join forums and online groups without the fear of dealing with the crypto shills
 I built FlipACoinFree.com — a fast, ad-free, embeddable coin-flip simulator with server-side randomness, API, and 30+ languages. Here’s how I built it.
I built FlipACoinFree.com — a fast, ad-free, embeddable coin-flip simulator with server-side randomness, API, and 30+ languages. Here’s how I built it.
 Discover why trying to please everyone in B2B product development leads to failure and how focusing on what matters can drive success in this 4-part series.
Discover why trying to please everyone in B2B product development leads to failure and how focusing on what matters can drive success in this 4-part series.
 *Note: This article first appeared in the Los Angeles Tribune by Hyro's Head of Marketing Aaron Bours
*Note: This article first appeared in the Los Angeles Tribune by Hyro's Head of Marketing Aaron Bours
 “In these unprecedented times…” People build unprecedented products, and contribute to the internet in unprecedented ways.
“In these unprecedented times…” People build unprecedented products, and contribute to the internet in unprecedented ways.
 Pursuing bold ideas at the right time and in the right place is the key to success in the world of entrepreneurship.
Pursuing bold ideas at the right time and in the right place is the key to success in the world of entrepreneurship.
 Explore why "some research" can harm innovation & sound design, plus tools & reads for better UX practices. Empower smarter research decisions today!
Explore why "some research" can harm innovation & sound design, plus tools & reads for better UX practices. Empower smarter research decisions today!
 There has been a huge influx of creativity in the innovation management software space. You must take a look at these differentiating features.
There has been a huge influx of creativity in the innovation management software space. You must take a look at these differentiating features.
 For people who are busy.
For people who are busy.
 Why the future belongs to those who think deeply — not those who write prettily.
Why the future belongs to those who think deeply — not those who write prettily.
 A look at what transpired at the 2nd Annual Innovate Summit, held in Owensboro, KY
A look at what transpired at the 2nd Annual Innovate Summit, held in Owensboro, KY
 One founder shares how he built an AI-powered executive team, including a GPT-based CTO, to drive strategy, creativity, and sustainable innovation.
One founder shares how he built an AI-powered executive team, including a GPT-based CTO, to drive strategy, creativity, and sustainable innovation.
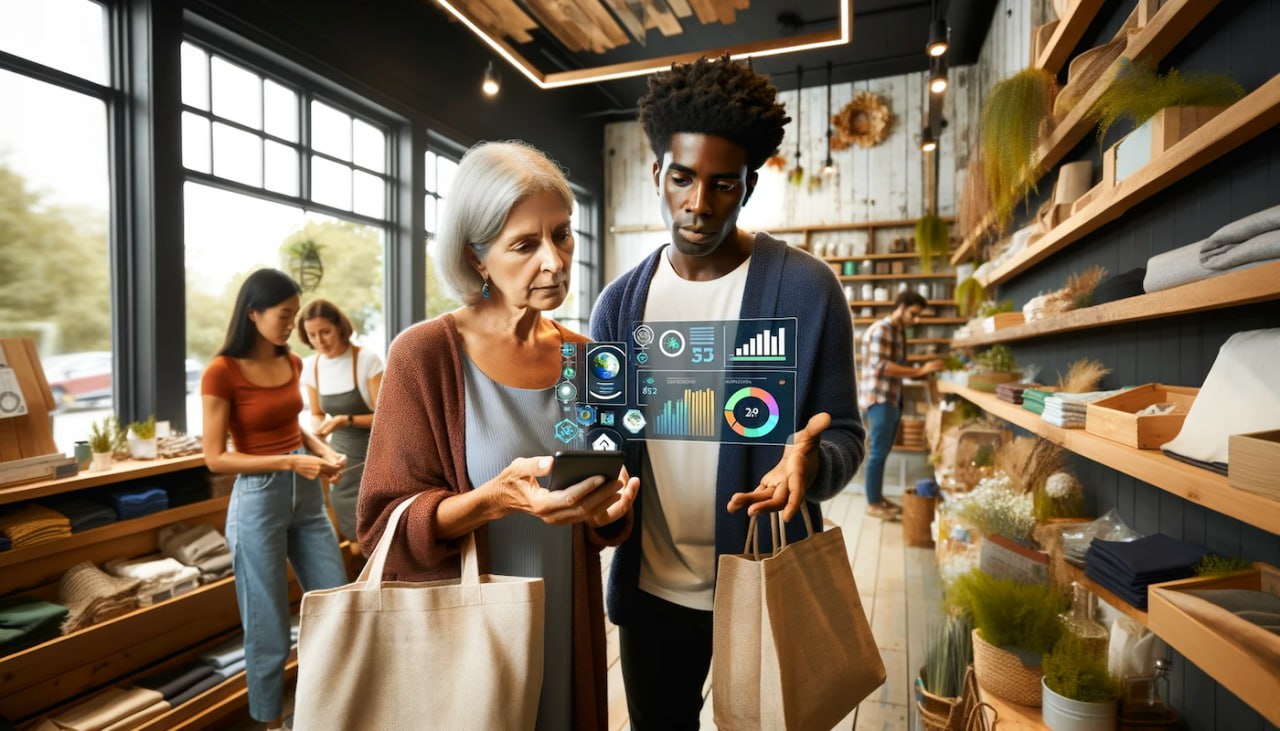 Discover how Impactful's CTO pioneers eco-friendly business analytics for a sustainable future in our insightful interview with Slav Dunaev.
Discover how Impactful's CTO pioneers eco-friendly business analytics for a sustainable future in our insightful interview with Slav Dunaev.
 TimeLens can understand the movement of the particles in-between the frames of a video to reconstruct what really happened at a speed even our eyes cannot see.
TimeLens can understand the movement of the particles in-between the frames of a video to reconstruct what really happened at a speed even our eyes cannot see.
 FOSS is the big boy LEGO of the Innovative World
FOSS is the big boy LEGO of the Innovative World
 Engineers are good at solutions. We're even fairly good at problems. Give us a client, and we will interview for the pain points, carefully describe the key problems, and design the most effective, doable solutions. And we'll do them, breaking them into actionable steps until the solution is realized.
Engineers are good at solutions. We're even fairly good at problems. Give us a client, and we will interview for the pain points, carefully describe the key problems, and design the most effective, doable solutions. And we'll do them, breaking them into actionable steps until the solution is realized.
 The "natural" world is not a sanctuary; it is a sequence of variables indifferent to your survival.
The "natural" world is not a sanctuary; it is a sequence of variables indifferent to your survival.
 Bernard Moon is Venture Capitalist, Co-Founder of SparkLabs group and 3x Noonies nominee.
Bernard Moon is Venture Capitalist, Co-Founder of SparkLabs group and 3x Noonies nominee.
 Juggling too many Proof of Concepts (POCs) and falling into the innovation buzzword trap can derail product development. Discover how to escape “Everyone” trap
Juggling too many Proof of Concepts (POCs) and falling into the innovation buzzword trap can derail product development. Discover how to escape “Everyone” trap
 This article series tries to put forward General Concepts, Principles and Guidelines for Innovative Software Product Design and Development
This article series tries to put forward General Concepts, Principles and Guidelines for Innovative Software Product Design and Development
 Throughout history, terrible crises have given us some of our greatest innovations. GPS, Drones, microwaves, atomic energy… the list goes on. The COVID-19 crisis is no different, and we are already seeing companies rise above their competition by asking themselves the key question at the heart of all corporate strategies : what can we do, right now, that is hard?
Throughout history, terrible crises have given us some of our greatest innovations. GPS, Drones, microwaves, atomic energy… the list goes on. The COVID-19 crisis is no different, and we are already seeing companies rise above their competition by asking themselves the key question at the heart of all corporate strategies : what can we do, right now, that is hard?
 Learn what IT leaders are prioritizing one year into the COVID-19 pandemic, per new research on the adoption of microservices, open-source tech, and more.
Learn what IT leaders are prioritizing one year into the COVID-19 pandemic, per new research on the adoption of microservices, open-source tech, and more.

 On June 4–5, Minsk (Belarus) hosted EMERGE, the largest IT conference in Eastern Europe. Speakers from all over the world flocked to the capital of one of the fastest growing technological hubs in the region. There, they shared their valuable insights on what’s on the tech table today. Iflexion lucked out to visit the conference and share its key takeaways with the world.
On June 4–5, Minsk (Belarus) hosted EMERGE, the largest IT conference in Eastern Europe. Speakers from all over the world flocked to the capital of one of the fastest growing technological hubs in the region. There, they shared their valuable insights on what’s on the tech table today. Iflexion lucked out to visit the conference and share its key takeaways with the world.
 Not all tech replaces the old. Many, like LLMs and quantum computers, create forks—parallel options, not revolutions. Knowing the difference helps us adapt.
Not all tech replaces the old. Many, like LLMs and quantum computers, create forks—parallel options, not revolutions. Knowing the difference helps us adapt.
 1/10/2025: Top 5 stories on the HackerNoon homepage!
1/10/2025: Top 5 stories on the HackerNoon homepage!
 Follow the story of Alex, a budding entrepreneur and discover the valuable ;lesson he learnt along the way.
Follow the story of Alex, a budding entrepreneur and discover the valuable ;lesson he learnt along the way.
 How innovation and development became the glue for Middle East peace.
How innovation and development became the glue for Middle East peace.
 To determine the relative smartness level of a city, there are several indicators that have been defined to arrive as such a number. These indicators, among many others, are:
To determine the relative smartness level of a city, there are several indicators that have been defined to arrive as such a number. These indicators, among many others, are:
 March 30, 1853. Vincent van Gogh was born. He is one of the most recognizable and influential painters of all time. But, what’s the link with human and machine, self-awareness and self-isolation? Well, something happened last week that made me question the way I think about our relationship with technology. And this started with Van Gogh.
March 30, 1853. Vincent van Gogh was born. He is one of the most recognizable and influential painters of all time. But, what’s the link with human and machine, self-awareness and self-isolation? Well, something happened last week that made me question the way I think about our relationship with technology. And this started with Van Gogh.
 Things have been a little fiery recently but since Elon is a cool dude I believe he can help Mother Nature chill out and not be so hot headed.
Things have been a little fiery recently but since Elon is a cool dude I believe he can help Mother Nature chill out and not be so hot headed.
 This is part of the reason that I launched the Open Innovation License prior. However, I felt like the OIN isn't enough. I felt like more needed to be done for the open source community and even the outside world that wants to establish a certain standard.
This is part of the reason that I launched the Open Innovation License prior. However, I felt like the OIN isn't enough. I felt like more needed to be done for the open source community and even the outside world that wants to establish a certain standard.

 Photo by Ameer Basheer on Unsplash
Photo by Ameer Basheer on Unsplash
If you’ve been to CES, you know it’s like a sprawling, exhausting rumpus room for tech geeks. You probably saw things you didn’t even believe were possible. And you probably went home overstimulated and mind-blown. But how does the technology extravaganza look to someone who knows tech inside and out, someone who keeps up with each new product as they’re released and developed?
 Businesses around the world have been forced to scramble a response to the current worldwide pandemic. Boardrooms have become (virtual) war rooms as organizations hunker down and try and devise strategies that will enable them to prosper or, in many cases, simply survive.
Businesses around the world have been forced to scramble a response to the current worldwide pandemic. Boardrooms have become (virtual) war rooms as organizations hunker down and try and devise strategies that will enable them to prosper or, in many cases, simply survive.
 In the last couples years, Mario Ribeiro Alves created Portugal's Blockchain Center, founded TAIKAI, AND got nominated for a 2020 Noonie for considered contributions to the INNOVATION category here at Hacker Noon.
In the last couples years, Mario Ribeiro Alves created Portugal's Blockchain Center, founded TAIKAI, AND got nominated for a 2020 Noonie for considered contributions to the INNOVATION category here at Hacker Noon.
 Fraud costs everyone: $6T globally, $500 per U.S. household. It’s the hidden tax draining wallets, businesses, and innovation—without you signing up.
Fraud costs everyone: $6T globally, $500 per U.S. household. It’s the hidden tax draining wallets, businesses, and innovation—without you signing up.
Product Development in a Lean and Efficient Way](https://hackernoon.com/lean-innovation-how-to-tackle-product-development-in-a-lean-and-efficient-way-tx2b3wqu)
 Nowadays, it is especially important to keep the process of product development lean and efficient. Therefore it is helpful to determine the concrete goal of each phase. This makes it easier to keep the entire process goal-oriented, structured and efficient.
Nowadays, it is especially important to keep the process of product development lean and efficient. Therefore it is helpful to determine the concrete goal of each phase. This makes it easier to keep the entire process goal-oriented, structured and efficient.
 Recent innovations in the construction industry include robotics and digital twins, enhancing productivity and efficiency.
Recent innovations in the construction industry include robotics and digital twins, enhancing productivity and efficiency.
 Fourth annual report reveals continued market maturation; $34.5 billion invested in southeastern startups since 2015
Fourth annual report reveals continued market maturation; $34.5 billion invested in southeastern startups since 2015
 Leading by example demonstrates a commitment to Innovation and risk-taking for results.
Leading by example demonstrates a commitment to Innovation and risk-taking for results.
 Founders chase speed, but forget the one metric that truly matters — validation velocity. Here’s why learning faster beats building faster in the AI era.
Founders chase speed, but forget the one metric that truly matters — validation velocity. Here’s why learning faster beats building faster in the AI era.
 United States Of America. v. Microsoft Corporation Court Filing by Thomas Penfield Jackson, November 5, 1999, is part of HackerNoon’s Legal PDF Series.
United States Of America. v. Microsoft Corporation Court Filing by Thomas Penfield Jackson, November 5, 1999, is part of HackerNoon’s Legal PDF Series.

 There has been a huge influx of creativity in the innovation management software space. You must take a look at these differentiating features.
There has been a huge influx of creativity in the innovation management software space. You must take a look at these differentiating features.
 Codacy has launched a fellowship program for open-source projects, encouraging OSS creators worldwide to apply to become “Codacy Pioneers.”
Codacy has launched a fellowship program for open-source projects, encouraging OSS creators worldwide to apply to become “Codacy Pioneers.”
 Information technology has had a high growth rate for years and there is a reason for that: a constant flow of innovations in technology, but also in business processes, as growing competition on the market has made innovation a must for every organisation. On the other side, the top skills missing among job applicants in the current world are problem solving, critical thinking, innovation and creativity.
Information technology has had a high growth rate for years and there is a reason for that: a constant flow of innovations in technology, but also in business processes, as growing competition on the market has made innovation a must for every organisation. On the other side, the top skills missing among job applicants in the current world are problem solving, critical thinking, innovation and creativity.
 TimeLens can understand the movement of the particles in-between the frames of a video to reconstruct what really happened at a speed even our eyes cannot see.
TimeLens can understand the movement of the particles in-between the frames of a video to reconstruct what really happened at a speed even our eyes cannot see.
Visit the /Learn Repo to find the most read blog posts about any technology.